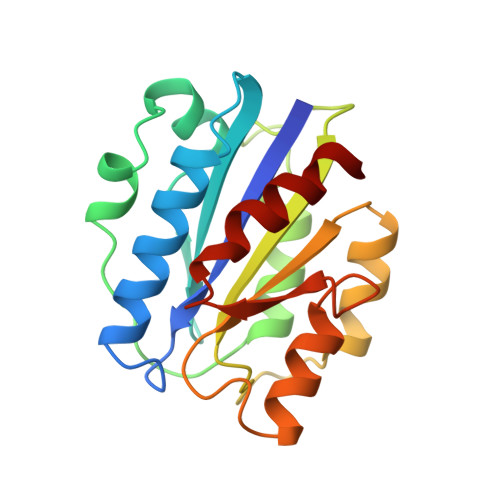
Crystal structure of the A domain from the alpha subunit of integrin CR3 (CD11b/CD18).
Lee, J.O., Rieu, P., Arnaout, M.A., Liddington, R.(1995) Cell 80: 631-638
- PubMed: 7867070 Search on PubMed
- DOI: https://doi.org/10.1016/0092-8674(95)90517-0
- Primary Citation Related Structures:
1IDO - PubMed Abstract:
We have determined the high resolution crystal structure of the A domain from the alpha chain of integrin CR3. The domain adopts a classic alpha/beta "Rossmann" fold and contains an unusual Mg2+ coordination site at its surface. One of the coordinating ligands is the glutamate side chain from another A domain molecule. We suggest that this site represents a general metal ion-dependent adhesion site (MIDAS) for binding protein ligands. We further propose that the beta subunits of integrins contain a MIDAS motif within a modified A domain. Our crystal structure will allow reliable models to be built for other members of the A domain superfamily and should facilitate development of novel adhesion modulatory drugs.
- Laboratory of X-Ray Crystallography, Dana Farber Cancer Institute, Harvard Medical School Boston, Massachusetts 02115.
Organizational Affiliation: