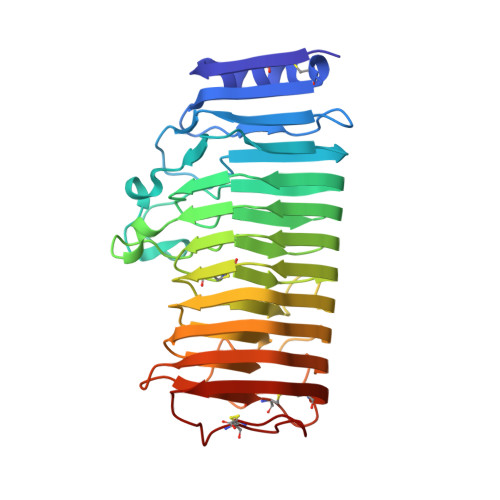
The X-ray structure of Aspergillus aculeatus polygalacturonase and a modeled structure of the polygalacturonase-octagalacturonate complex.
Cho, S.W., Lee, S., Shin, W.(2001) J Mol Biology 311: 863-878
- PubMed: 11518536 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4919
- Primary Citation Related Structures:
1IA5, 1IB4 - PubMed Abstract:
Polygalacturonases hydrolyze the alpha-(1-4) glycosidic bonds of de-esterified pectate in the smooth region of the plant cell wall. Crystal structures of polygalacturonase from Aspergillus aculeatus were determined at pH 4.5 and 8.5 both to 2.0 A resolution. A. aculeatus polygalacturonase is a glycoprotein with one N and ten O-glycosylation sites and folds into a right-handed parallel beta-helix. The structures of the three independent molecules are essentially the same, showing no dependency on pH or crystal packing, and are very similar to that of Aspergillus niger polygalacturonase. However, the structures of the long T1 loop containing a catalytic tyrosine residue are significantly different in the two proteins. A three-dimensional model showing the substrate binding mode for a family 28 hydrolase was obtained by a combined approach of flexible docking, molecular dynamics simulations, and energy minimization. The octagalacturonate substrate was modeled as an unbent irregular helix with the -1 ring in a half-chair ((4)H(3)) form that approaches the transition state conformation. A comparative modeling of the three polygalacturonases with known structure shows that six subsites ranging from -4 to +2 are clearly defined but subsites -5 and +3 may or may not be shaped depending on the nearby amino acid residues. Both distal subsites are mostly exposed to the solvent region and have weak binding affinity even if they exist. The complex model provides a clear explanation for the functions, either in catalysis or in substrate binding, of all conserved amino acid residues in the polygalacturonase family of proteins. Modeling suggests that the role of the conserved Asn157 and Tyr270, which had previously been unidentified, may be in transition state stabilization. In A. niger polygalacturonase, the long T1 loop may have to undergo conformational change upon binding of the substrate to bring the tyrosine residue close to subsite -1.
- School of Chemistry and Molecular Engineering, and Center for Molecular Catalysis, Seoul National University, Seoul, 151-742, Korea.
Organizational Affiliation: