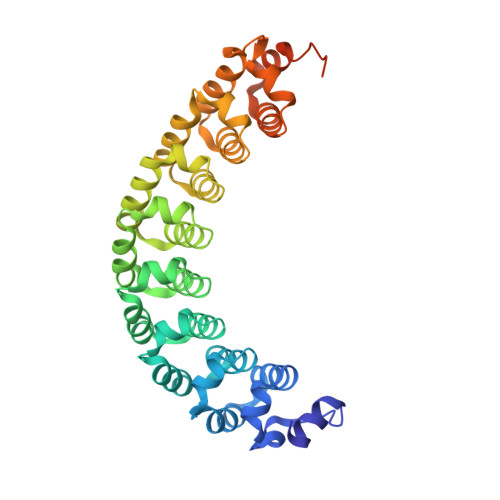
Crystal structure of a Pumilio homology domain.
Wang, X., Zamore, P.D., Hall, T.M.(2001) Mol Cell 7: 855-865
- PubMed: 11336708 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(01)00229-5
- Primary Citation Related Structures:
1IB2, 1M8Z - PubMed Abstract:
Puf proteins regulate translation and mRNA stability by binding sequences in their target RNAs through the Pumilio homology domain (PUM-HD), which is characterized by eight tandem copies of a 36 amino acid motif, the PUM repeat. We have solved the structure of the PUM-HD from human Pumilio1 at 1.9 A resolution. The structure reveals that the eight PUM repeats correspond to eight copies of a single, repeated structural motif. The PUM repeats pack together to form a right-handed superhelix that approximates a half doughnut. The distribution of side chains on the inner and outer faces of this half doughnut suggests that the inner face of the PUM-HD binds RNA while the outer face interacts with proteins such as Nanos, Brain Tumor, and cytoplasmic polyadenylation element binding protein.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, NC 27709, USA.
Organizational Affiliation: