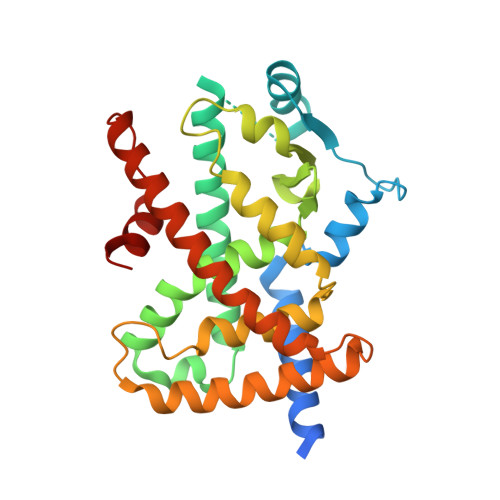
Structure of the PPARalpha and -gamma ligand binding domain in complex with AZ 242; ligand selectivity and agonist activation in the PPAR family.
Cronet, P., Petersen, J.F., Folmer, R., Blomberg, N., Sjoblom, K., Karlsson, U., Lindstedt, E.L., Bamberg, K.(2001) Structure 9: 699-701
- PubMed: 11587644 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00634-7
- Primary Citation Related Structures:
1I7G, 1I7I - PubMed Abstract:
The peroxisome proliferator-activated receptors (PPAR) are ligand-activated transcription factors belonging to the nuclear receptor family. The roles of PPARalpha in fatty acid oxidation and PPARgamma in adipocyte differentiation and lipid storage have been characterized extensively. PPARs are activated by fatty acids and eicosanoids and are also targets for antidyslipidemic drugs, but the molecular interactions governing ligand selectivity for specific subtypes are unclear due to the lack of a PPARalpha ligand binding domain structure. We have solved the crystal structure of the PPARalpha ligand binding domain (LBD) in complex with the combined PPARalpha and -gamma agonist AZ 242, a novel dihydro cinnamate derivative that is structurally different from thiazolidinediones. In addition, we present the crystal structure of the PPARgamma_LBD/AZ 242 complex and provide a rationale for ligand selectivity toward the PPARalpha and -gamma subtypes. Heteronuclear NMR data on PPARalpha in both the apo form and in complex with AZ 242 shows an overall stabilization of the LBD upon agonist binding. A comparison of the novel PPARalpha/AZ 242 complex with the PPARgamma/AZ 242 complex and previously solved PPARgamma structures reveals a conserved hydrogen bonding network between agonists and the AF2 helix. The complex of PPARalpha and PPARgamma with the dual specificity agonist AZ 242 highlights the conserved interactions required for receptor activation. Together with the NMR data, this suggests a general model for ligand activation in the PPAR family. A comparison of the ligand binding sites reveals a molecular explanation for subtype selectivity and provides a basis for rational drug design.
- Department of Molecular Biology, AstraZeneca R&D Mölndal, S-431 83, Mölndal, Sweden.
Organizational Affiliation: