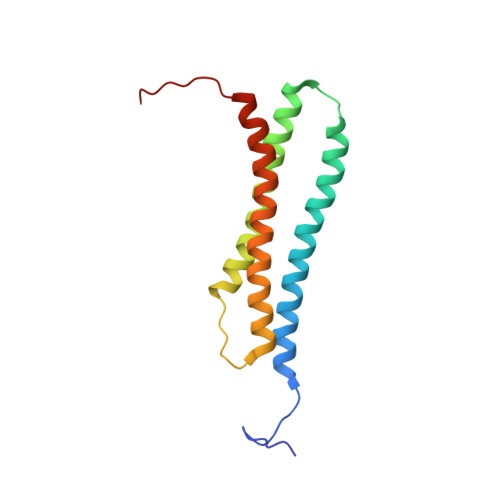
Structural analysis of BAG1 cochaperone and its interactions with Hsc70 heat shock protein.
Briknarova, K., Takayama, S., Brive, L., Havert, M.L., Knee, D.A., Velasco, J., Homma, S., Cabezas, E., Stuart, J., Hoyt, D.W., Satterthwait, A.C., Llinas, M., Reed, J.C., Ely, K.R.(2001) Nat Struct Biol 8: 349-352
- PubMed: 11276257 Search on PubMed
- DOI: https://doi.org/10.1038/86236
- Primary Citation Related Structures:
1I6Z - PubMed Abstract:
BAG-family proteins share a conserved protein interaction region, called the 'BAG domain', which binds and regulates Hsp70/Hsc70 molecular chaperones. This family of cochaperones functionally regulates signal transducing proteins and transcription factors important for cell stress responses, apoptosis, proliferation, cell migration and hormone action. Aberrant overexpression of the founding member of this family, BAG1, occurs in human cancers. In this study, a structure-based approach was used to identify interacting residues in a BAG1--Hsc70 complex. An Hsc70-binding fragment of BAG1 was shown by multidimensional NMR methods to consist of an antiparallel three-helix bundle. NMR chemical shift experiments marked surface residues on the second (alpha 2) and third (alpha 3) helices in the BAG domain that are involved in chaperone binding. Structural predictions were confirmed by site-directed mutagenesis of these residues, resulting in loss of binding of BAG1 to Hsc70 in vitro and in cells. Molecular docking of BAG1 to Hsc70 and mutagenesis of Hsc70 marked the molecular surface of the ATPase domain necessary for interaction with BAG1. The results provide a structural basis for understanding the mechanism by which BAG proteins link molecular chaperones and cell signaling pathways.
- The Burnham Institute, La Jolla, California 92037, USA.
Organizational Affiliation: