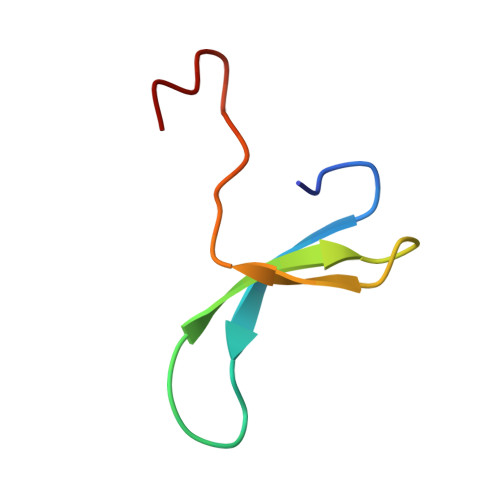
1H NMR study on the binding of Pin1 Trp-Trp domain with phosphothreonine peptides.
Wintjens, R., Wieruszeski, J.M., Drobecq, H., Rousselot-Pailley, P., Buee, L., Lippens, G., Landrieu, I.(2001) J Biological Chem 276: 25150-25156
- PubMed: 11313338
- DOI: https://doi.org/10.1074/jbc.M010327200
- Primary Citation Related Structures:
1I6C, 1I8G, 1I8H - PubMed Abstract:
The recent crystal structure of Pin1 protein bound to a doubly phosphorylated peptide from the C-terminal domain of RNA polymerase II revealed that binding interactions between Pin1 and its substrate take place through its Trp-Trp (WW) domain at the level of the loop Ser(11)-Arg(12) and the aromatic pair Tyr(18)-Trp(29), and showed a trans conformation for both pSer-Pro peptide bonds. However, the orientation of the ligand in the aromatic recognition groove still could be sequence-specific, as previously observed in SH3 domains complexed by peptide ligands or for different class of WW domains (Zarrinpar, A., and Lim, W. A. (2000) Nat. Struct. Biol. 7, 611-613). Because the bound peptide conformation could also differ as observed for peptide ligands bound to the 14-3-3 domain, ligand orientation and conformation for two other biologically relevant monophosphate substrates, one derived from the Cdc25 phosphatase of Xenopus laevis (EQPLpTPVTDL) and another from the human tau protein (KVSVVRpTPPKSPS) in complex with the WW domain are here studied by solution NMR methods. First, the proton resonance perturbations on the WW domain upon complexation with both peptide ligands were determined to be essentially located in the positively charged beta-hairpin Ser(11)-Gly(15) and around the aromatic Trp(29). Dissociation equilibrium constants of 117 and 230 microm for Cdc25 and tau peptides, respectively, were found. Several intermolecular nuclear Overhauser effects between WW domain and substrates were obtained from a ligand-saturated solution and were used to determine the structures of the complexes in solution. We found a similar N to C orientation as the one observed in the crystal complex structure of Pin1 and a trans conformation for the pThr-Pro peptidic bond in both peptide ligands, thereby indicating a unique binding scheme for the Pin1 WW domain to its multiple substrates.
- Institut de Biologie de Lille, Institut Pasteur de Lille, CNRS-UMR 8525, 1 rue du Professeur Calmette, BP 447, Lille 59021, France.
Organizational Affiliation: