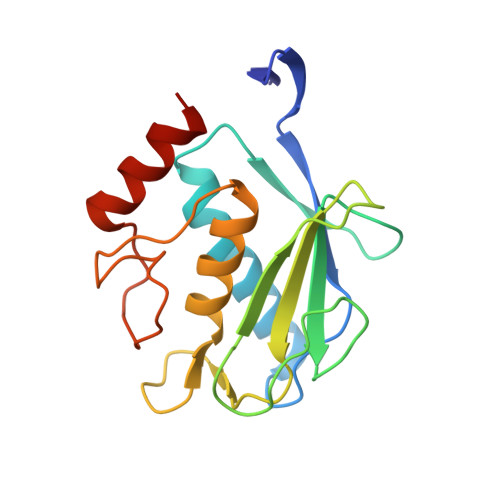
Crystal structure of the stromelysin-3 (MMP-11) catalytic domain complexed with a phosphinic inhibitor mimicking the transition-state.
Gall, A.L., Ruff, M., Kannan, R., Cuniasse, P., Yiotakis, A., Dive, V., Rio, M.C., Basset, P., Moras, D.(2001) J Mol Biology 307: 577-586
- PubMed: 11254383 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4493
- Primary Citation Related Structures:
1HV5 - PubMed Abstract:
Stromelysin-3 (ST3) is a matrix metalloproteinase (MMP-11) whose proteolytic activity plays an important role in tumorigenicity enhancement. In breast cancer, ST3 is a bad prognosis marker: its expression is associated with a poor clinical outcome. This enzyme therefore represents an attractive therapeutic target. The topology of matrix metalloproteinases (MMPs) is remarkably well conserved, making the design of highly specific inhibitors difficult. The major difference between MMPs lies in the S(1)' subsite, a well-defined hydrophobic pocket of variable depth. The present crystal structure, the first 3D-structure of the ST3 catalytic domain in interaction with a phosphinic inhibitor mimicking a (d, l) peptide, clearly demonstrates that its S(1)' pocket corresponds to a tunnel running through the enzyme. This open channel is filled by the inhibitor P(1)' group which adopts a constrained conformation to fit this pocket, together with two water molecules interacting with the ST3-specific residue Gln215. These observations provide clues for the design of more specific inhibitors and show how ST3 can accommodate a phosphinic inhibitor mimicking a (d, l) peptide. The presence of a water molecule interacting with one oxygen atom of the inhibitor phosphinyl group and the proline residue of the Met-turn suggests how the intermediate formed during proteolysis may be stabilized. Furthermore, the hydrogen bond distance observed between the methyl of the phosphinic group and the carbonyl group of Ala182 mimics the interaction between this carbonyl group and the amide group of the cleaved peptidic bond. Our crystal structure provides a good model to study the MMPs mechanism of proteolysis.
- Structural Biology and Genomics Laboratory, I.G.B.M.C., B.P. 163, F67404, Illkirch Cedex, France.
Organizational Affiliation: