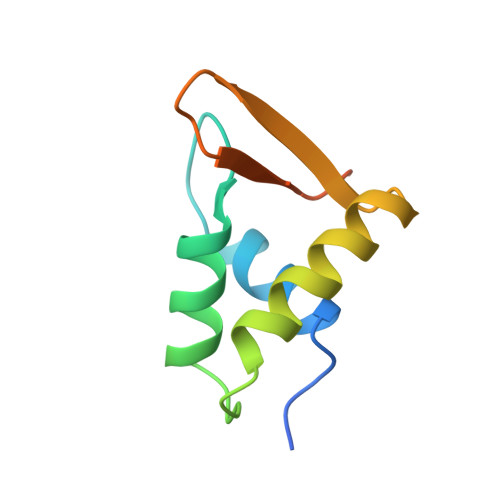
Crystal structure of globular domain of histone H5 and its implications for nucleosome binding.
Ramakrishnan, V., Finch, J.T., Graziano, V., Lee, P.L., Sweet, R.M.(1993) Nature 362: 219-223
- PubMed: 8384699 Search on PubMed
- DOI: https://doi.org/10.1038/362219a0
- Primary Citation Related Structures:
1HST - PubMed Abstract:
The structure of GH5, the globular domain of the linker histone H5, has been solved to 2.5 A resolution by multiwavelength anomalous diffraction on crystals of the selenomethionyl protein. The structure shows a striking similarity to the DNA-binding domain of the catabolite gene activator protein CAP, thereby providing a possible model for the binding of GH5 to DNA.
- Biology Department, Brookhaven National Laboratory, Upton, New York 11973.
Organizational Affiliation: