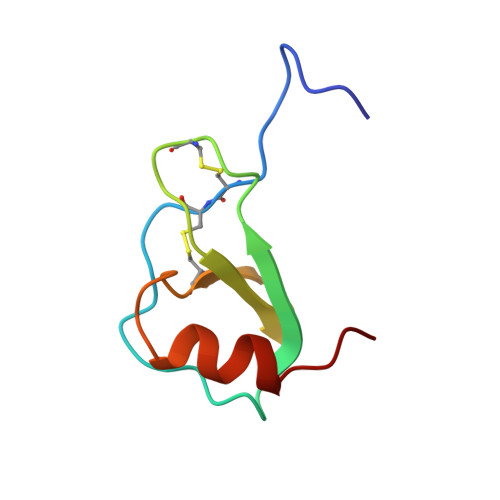
The three-dimensional solution structure of RANTES.
Chung, C.W., Cooke, R.M., Proudfoot, A.E., Wells, T.N.(1995) Biochemistry 34: 9307-9314
- PubMed: 7542919 Search on PubMed
- DOI: https://doi.org/10.1021/bi00029a005
- Primary Citation Related Structures:
1HRJ - PubMed Abstract:
The solution structure of the chemokine RANTES (regulated on activation, normal T-cell expressed and secreted) has been determined using NMR spectroscopy. Backbone and side-chain 1H and 15N assignments have been obtained using a combination of two-dimensional homonuclear and three-dimensional heteronuclear spectra. Regular elements of secondary structure have been identified on the basis of a qualitative interpretation of NOE data, J(NH-H alpha) coupling constants, and amide exchange rates. Three-dimensional structures were calculated from a total of 2146 experimental restraints using a combination of distance geometry and simulated annealing protocols. For the 13 best structures the average backbone (N, C alpha, C) atomic rmsd from the mean coordinates for residues 5-65 is 0.64 A (+/- 0.14 A) for the dimer and 0.50 A (+/- 0.08 A) for the individual monomers. Each monomer consists of a three-stranded antiparallel beta-sheet (residues 26-30, 38-43, 48-51) in a Greek key motif with a C-terminal helix (56-65) packed across the sheet, an arrangement similar to the monomeric structure of other members of this chemokine family (IL-8, PF4, MGSA/Gro alpha, and MIP-1 beta). Overall, the RANTES dimer resembles that previously reported for MIP-1 beta.
- Department of Biomolecular Structure, Glaxo Research and Development Ltd., Hertfordshire, U.K.
Organizational Affiliation: