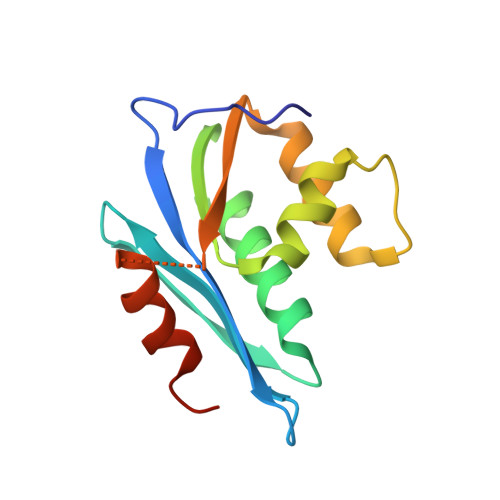
Crystal structure of the ribonuclease H domain of HIV-1 reverse transcriptase.
Davies 2nd., J.F., Hostomska, Z., Hostomsky, Z., Jordan, S.R., Matthews, D.A.(1991) Science 252: 88-95
- PubMed: 1707186
- DOI: https://doi.org/10.1126/science.1707186
- Primary Citation Related Structures:
1HRH - PubMed Abstract:
The crystal structure of the ribonuclease (RNase) H domain of HIV-1 reverse transcriptase (RT) has been determined at a resolution of 2.4 A and refined to a crystallographic R factor of 0.20. The protein folds into a five-stranded mixed beta sheet flanked by an asymmetric distribution of four alpha helices. Two divalent metal cations bind in the active site surrounded by a cluster of four conserved acidic amino acid residues. The overall structure is similar in most respects to the RNase H from Escherichia coli. Structural features characteristic of the retroviral protein suggest how it may interface with the DNA polymerase domain of p66 in the mature RT heterodimer. These features also offer insights into why the isolated RNase H domain is catalytically inactive but when combined in vitro with the isolated p51 domain of RT RNase H activity can be reconstituted. Surprisingly, the peptide bond cleaved by HIV-1 protease near the polymerase-RNase H junction of p66 is completely inaccessible to solvent in the structure reported here. This suggests that the homodimeric p66-p66 precursor of mature RT is asymmetric with one of the two RNase H domains at least partially unfolded.
- Agouron Pharmaceuticals, Inc., La Jolla, CA 92037.
Organizational Affiliation: