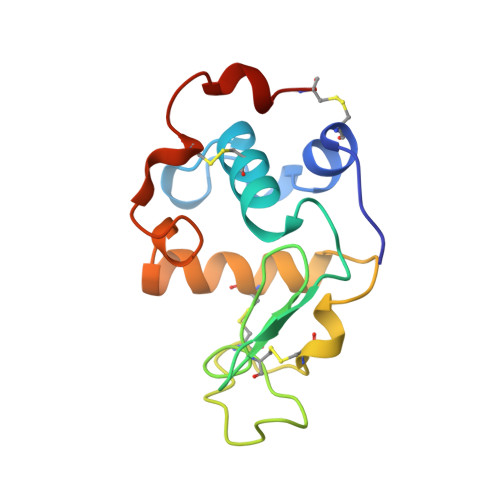
Effect of the extra n-terminal methionine residue on the stability and folding of recombinant alpha-lactalbumin expressed in Escherichia coli.
Chaudhuri, T.K., Horii, K., Yoda, T., Arai, M., Nagata, S., Terada, T.P., Uchiyama, H., Ikura, T., Tsumoto, K., Kataoka, H., Matsushima, M., Kuwajima, K., Kumagai, I.(1999) J Mol Biology 285: 1179-1194
- PubMed: 9887272
- DOI: https://doi.org/10.1006/jmbi.1998.2362
- Primary Citation of Related Structures:
1HMK - PubMed Abstract:
The structure, stability, and unfolding-refolding kinetics of Escherichia coli-expressed recombinant goat alpha-lactalbumin were studied by circular dichroism spectroscopy, X-ray crystallography, and stopped-flow measurements, and the results were compared with those of the authentic protein prepared from goat milk. The electric properties of the two proteins were also studied by gel electrophoresis and ion-exchange chromatography. Although the overall structures of the authentic and recombinant proteins are the same, the extra methionine residue at the N terminus of the recombinant protein remarkably affects the native-state stability and the electric properties. The native state of the recombinant protein was 3.5 kcal/mol less stable than the authentic protein, and the recombinant protein was more negatively charged than the authentic one. The recombinant protein unfolded 5.7 times faster than the authentic one, although there were no significant differences in the refolding rates of the two proteins. The destabilization of the recombinant protein can be fully interpreted in terms of the increased unfolding rate of the protein, indicating that the N-terminal region remains unorganized in the transition state of refolding, and hence is not involved in the folding initiation site of the protein. A comparison of the X-ray structures of recombinant alpha-lactalbumin determined here with that of the authentic protein shows that the structural differences between the proteins are confined to the N-terminal region. Theoretical considerations for the differences in the conformational and solvation free energies between the proteins show that the destabilization of the recombinant protein is primarily due to excess conformational entropy of the N-terminal methionine residue in the unfolded state, and also due to less exposure of hydrophobic surface on unfolding. The results suggest that when the N-terminal region of a protein has a rigid structure, expression of the protein by E. coli, which adds the extra methionine residue, destabilizes the native state through a conformational entropy effect. It also shows that differences in the electrostatic interactions of the N-terminal amino group with the side-chain atoms of Thr38, Asp37, and Asp83 bring about a difference in the pKa value of the N-terminal amino group between the proteins, resulting in a greater negative net charge of the recombinant protein at neutral pH.
- Department of Physics Graduate School of Science, University of Tokyo, Tokyo, 113-0033, Japan.
Organizational Affiliation: