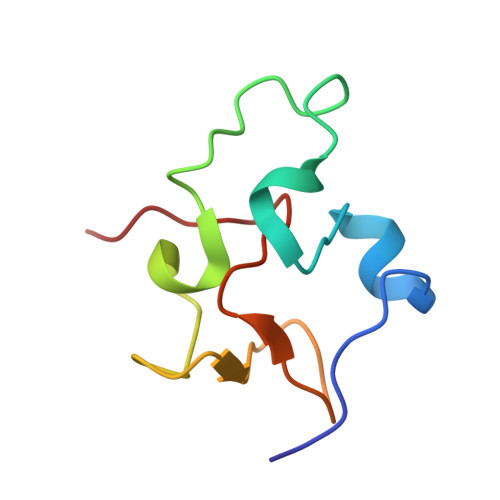
Structure of Rhodoferax fermentans high-potential iron-sulfur protein solved by MAD.
Gonzalez, A., Benini, S., Ciurli, S.(2003) Acta Crystallogr D Biol Crystallogr 59: 1582-1588
- PubMed: 12925788
- DOI: https://doi.org/10.1107/s0907444903014604
- Primary Citation of Related Structures:
1HLQ - PubMed Abstract:
The crystal structure of Rhodoferax fermentans high-potential iron protein (HiPIP) has been solved by MAD methods using the anomalous signal from the Fe atoms in the [Fe(4)S(4)] cluster present in the protein and refined to a resolution of 1.45 A. The peptide chain is well defined except in the N- and C-terminal areas. The structure of the protein reveals the presence of three helical fragments, a small beta-sheet and several turns, with the [Fe(4)S(4)] cluster being located close to a surface patch containing several well conserved aromatic residues. The protein fold is very similar to the structures of other known HiPIPs, especially in the region proximal to the [Fe(4)S(4)] cluster, while the largest differences are observed on the opposite side of the protein, which is rich in positive charges and has no sequential homology to other HiPIP families.
- Stanford Synchrotron Radiation Laboratory, 2575 Sand Hill Road MS99, Menlo Park, CA 94025, USA.
Organizational Affiliation: