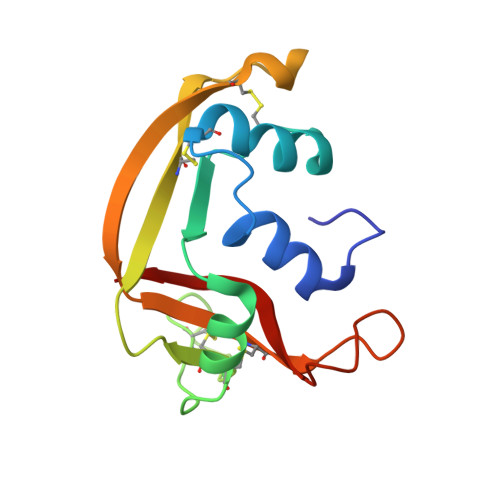
Mapping the Ribonucleolytic Active Site of Eosinophil-Derived Neurotoxin (Edn): High Resolution Crystal Structures of Edn Complexes with Adenylic Nucleotide Inhibitors
Leonidas, D.D., Boix, E., Prill, R., Suzuki, M., Turton, R., Minson, K., Swaminathan, G.J., Youle, R.J., Acharya, K.R.(2001) J Biological Chem 276: 15009
- PubMed: 11154698 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M010585200
- Primary Citation Related Structures:
1HI2, 1HI3, 1HI4, 1HI5 - PubMed Abstract:
Eosinophil-derived neurotoxin (EDN), a basic ribonuclease found in the large specific granules of eosinophils, belongs to the pancreatic RNase A family. Although its physiological function is still unclear, it has been shown that EDN is a neurotoxin capable of inducing the Gordon phenomenon in rabbits. EDN is also a potent helminthotoxin and can mediate antiviral activity of eosinophils against isolated virions of the respiratory syncytial virus. EDN is a catalytically efficient RNase sharing similar substrate specificity with pancreatic RNase A with its ribonucleolytic activity being absolutely essential for its neurotoxic, helminthotoxic, and antiviral activities. The crystal structure of recombinant human EDN in the unliganded form has been determined previously (Mosimann, S. C., Newton, D. L., Youle, R. J., and James, M. N. G. (1996) J. Mol. Biol. 260, 540-552). We have now determined high resolution (1.8 A) crystal structures for EDN in complex with adenosine-3',5'-diphosphate (3',5'-ADP), adenosine-2',5'-di-phosphate (2',5'-ADP), adenosine-5'-diphosphate (5'-ADP) as well as for a native structure in the presence of sulfate refined at 1.6 A. The inhibition constant of these mononucleotides for EDN has been determined. The structures present the first detailed picture of differences between EDN and RNase A in substrate recognition at the ribonucleolytic active site. They also provide a starting point for the design of tight-binding inhibitors, which may be used to restrain the RNase activity of EDN.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: