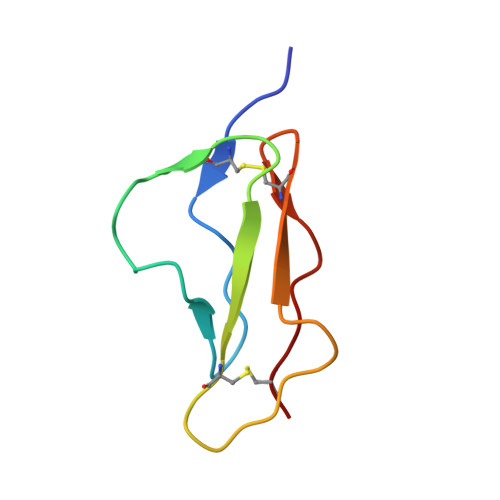
Solution structure of a pair of complement modules by nuclear magnetic resonance.
Barlow, P.N., Steinkasserer, A., Norman, D.G., Kieffer, B., Wiles, A.P., Sim, R.B., Campbell, I.D.(1993) J Mol Biology 232: 268-284
- PubMed: 8331663 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1381
- Primary Citation Related Structures:
1HFH, 1HFI - PubMed Abstract:
A portion of human complement factor H spanning the 15th (H15) and 16th (H16) of its 20 modules, has been expressed in a yeast vector and subjected to structure determination in solution using two-dimensional 1H-NMR. The structure of H15 is very similar to that already established for the fifth module of factor H and H16, consistent with the view that all such complement control (C-) modules share a common overall topology. In addition, the tertiary structures of the component modules of the H15-16 pair are very similar to those of the modules when expressed individually, implying that each folds entirely autonomously within intact factor H. Aromatic residues in the third turn of H15 and the second turn of H16, together with a leucine residue from the linker region, contribute to a small intermodular interface. Comparatively few nuclear Overhauser effects were observable between protons on different modules. Consequently, a wide range of angles of "twist" (131 (+/- 146) degrees, mean value (+/- 1 standard deviation)), i.e. rotation about the long axis of one module with respect to the other, exists in the family of structures generated on the basis of the experimental data. However, much smaller variations occur in the two, orthogonal, angles (175 (+/- 12) degrees and 103 (+/- 6) degrees) that describe the "tilt". These observations may suggest upper limits on the relative flexibility of the two modules. Models were built to assess the outcome of applying such restrictions to all the neighbours within a string of 20 C-modules, and the resulting structures compare well with factor H as visualized by electron microscopy.
- Department of Biochemistry, University of Oxford, U.K.
Organizational Affiliation: