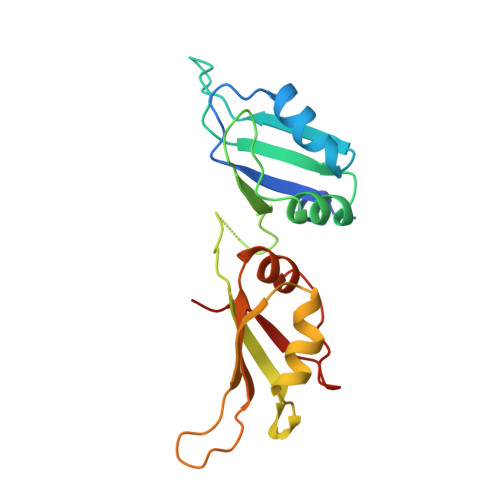
Crystal structure of the two RNA binding domains of human hnRNP A1 at 1.75 A resolution.
Shamoo, Y., Krueger, U., Rice, L.M., Williams, K.R., Steitz, T.A.(1997) Nat Struct Biol 4: 215-222
- PubMed: 9164463 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0397-215
- Primary Citation Related Structures:
1HA1 - PubMed Abstract:
Heterogeneous ribonucleoprotein A1 (hnRNP A1) is an abundant eukaryotic nuclear RNA binding protein. A1 is involved in the packaging of pre-mRNA into hnRNP particles, transport of poly A+ mRNA from the nucleus to the cytoplasm and may modulate splice site selection. The crystal structure of A1(RBD1,2) reveals two independently-folded RNA binding domains (RBDs) connected by a flexible linker. Both RBDs are structurally homologous to the U1A(RBD1), and have their RNA binding platforms oriented in an anti-parallel fashion. The anti-parallel arrangement of the A1 RNA binding platforms suggests mechanisms for RNA condensation and ways of bringing together distant RNA sequences for RNA metabolism such as splicing or transport.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06520-8114, USA.
Organizational Affiliation: