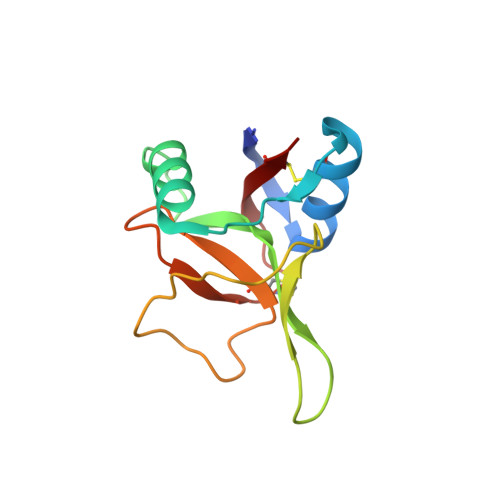
Crystal Structure of the Eosinophil Major Basic Protein at 1.8A. An Atypical Lectin with a Paradigm Shift in Specificity
Swaminathan, G.J., Weaver, A.J., Loegering, D.A., Checkel, J.L., Leonidas, D.D., Gleich, G.J., Acharya, K.R.(2001) J Biological Chem 276: 26197
- PubMed: 11319227 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M100848200
- Primary Citation Related Structures:
1H8U - PubMed Abstract:
The eosinophil major basic protein (EMBP) is the predominant constituent of the crystalline core of the eosinophil primary granule. EMBP is directly implicated in epithelial cell damage, exfoliation, and bronchospasm in allergic diseases such as asthma. Here we report the crystal structure of EMBP at 1.8 A resolution, and show that it is similar to that of members of the C-type lectin superfamily with which it shares minimal amino acid sequence identity (approximately 15--28%). However, this protein lacks a Ca(2+)/carbohydrate-binding site. Our analysis suggests that EMBP specifically binds heparin. Based on our results, we propose a possible new function for this protein, which is likely to have implications for EMBP function.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: