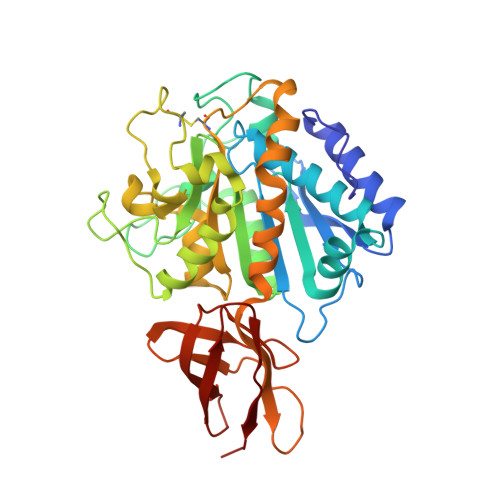
The crystal structure of the inhibitor-complexed carboxypeptidase D domain II and the modeling of regulatory carboxypeptidases.
Aloy, P., Companys, V., Vendrell, J., Aviles, F.X., Fricker, L.D., Coll, M., Gomis-Ruth, F.X.(2001) J Biological Chem 276: 16177-16184
- PubMed: 11278909 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M011457200
- Primary Citation Related Structures:
1H8L - PubMed Abstract:
The three-dimensional crystal structure of duck carboxypeptidase D domain II has been solved in a complex with the peptidomimetic inhibitor, guanidinoethylmercaptosuccinic acid, occupying the specificity pocket. This structure allows a clear definition of the substrate binding sites and the substrate funnel-like access. The structure of domain II is the only one available from the regulatory carboxypeptidase family and can be used as a general template for its members. Here, it has been used to model the structures of domains I and III from the former protein and of human carboxypeptidase E. The models obtained show that the overall topology is similar in all cases, the main differences being local and because of insertions in non-regular loops. In both carboxypeptidase D domain I and carboxypeptidase E slightly different shapes of the access to the active site are predicted, implying some kind of structural selection of protein or peptide substrates. Furthermore, emplacement of the inhibitor structure in the active site of the constructed models showed that the inhibitor fits very well in all of them and that the relevant interactions observed with domain II are conserved in domain I and carboxypeptidase E but not in the non-active domain III because of the absence of catalytically indispensable residues in the latter protein. However, in domain III some of the residues potentially involved in substrate binding are well preserved, together with others of unknown roles, which also are highly conserved among all carboxypeptidases. These observations, taken together with others, suggest that domain III might play a role in the binding and presentation of proteins or peptide substrates, such as the pre-S domain of the large envelope protein of duck hepatitis B virus.
- Institut de Biologia Fonamental and Departament de Bioquimica i Biologia Molecular, Unitat de Ciències, Universitat Autònoma de Barcelona, 08193 Bellaterra, Barcelona, Spain.
Organizational Affiliation: