Mechanism of C-Myb-C/Ebpbeta Cooperation from Separated Sites on a Promoter
Tahirov, T.H., Sato, K., Ichikawa-Iwata, E., Sasaki, M., Inoue-Bungo, T., Shiina, M., Kimura, K., Takata, S., Fujikawa, A., Morii, H., Kumasaka, T., Yamamoto, M., Ishii, S., Ogata, K.(2002) Cell 108: 57
- PubMed: 11792321 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(01)00636-5
- Primary Citation Related Structures:
1H88, 1H89, 1H8A - PubMed Abstract:
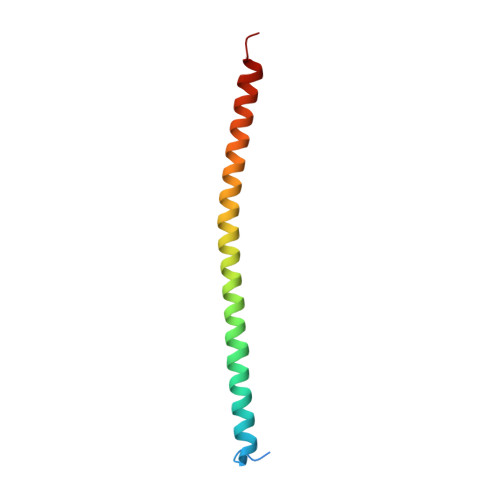
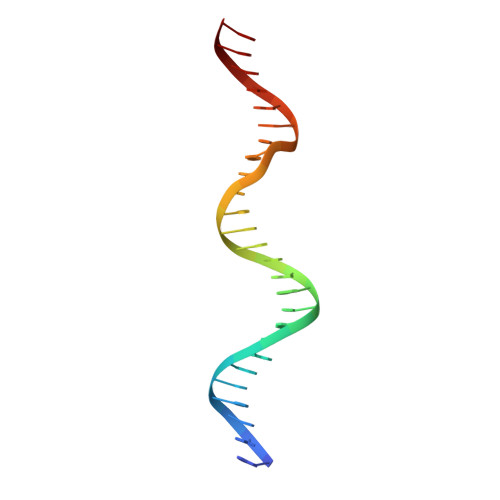
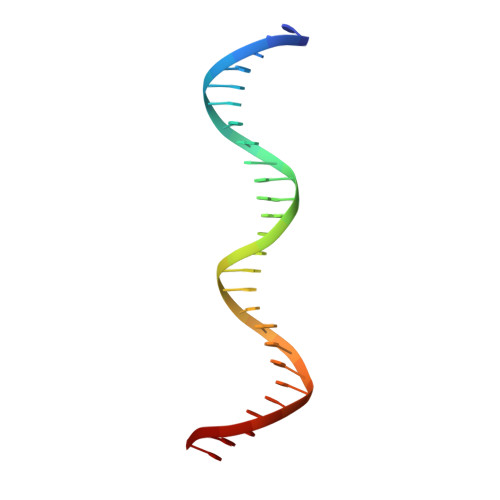
c-Myb, but not avian myeloblastosis virus (AMV) v-Myb, cooperates with C/EBP beta to regulate transcription of myeloid-specific genes. To assess the structural basis for that difference, we determined the crystal structures of complexes comprised of the c-Myb or AMV v-Myb DNA-binding domain (DBD), the C/EBP beta DBD, and a promoter DNA fragment. Within the c-Myb complex, a DNA-bound C/EBP beta interacts with R2 of c-Myb bound to a different DNA fragment; point mutations in v-Myb R2 eliminate such interaction within the v-Myb complex. GST pull-down assays, luciferase trans-activation assays, and atomic force microscopy confirmed that the interaction of c-Myb and C/EBP beta observed in crystal mimics their long range interaction on the promoter, which is accompanied by intervening DNA looping.
- Kanagawa Academy of Science and Technology, 3-9 Fukuura, Kanazawa-ku, Yokohama 236-0004, Japan. tahir@med.yokohama-cu.ac.jp
Organizational Affiliation: