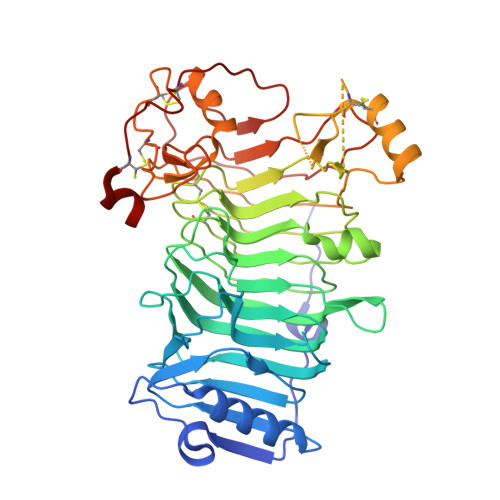
The Iota-Carrageenase of Alteromonas Fortis. A Beta-Helix Fold-Containing Enzyme for the Degradation of a Highly Polyanionic Polysaccharide
Michel, G., Chantalat, L., Fanchon, E., Henrissat, B., Kloareg, B., Dideberg, O.(2001) J Biological Chem 276: 40202
- PubMed: 11493601 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M100670200
- Primary Citation Related Structures:
1H80 - PubMed Abstract:
Carrageenans are gel-forming hydrocolloids extracted from the cell walls of marine red algae. They consist of d-galactose residues bound by alternate alpha(1-->3) and beta(1-->4) linkages and substituted by one (kappa-carrageenan), two (iota-carrageenan), or three (lambda-carrageenan) sulfate-ester groups per disaccharide repeating unit. Both the kappa- and iota-carrageenan chains adopt ordered conformations leading to the formation of highly ordered aggregates of double-stranded helices. Several kappa-carrageenases and iota-carrageenases have been cloned from marine bacteria. Kappa-carrageenases belong to family 16 of the glycoside hydrolases, which essentially encompasses polysaccharidases specialized in the hydrolysis of the neutral polysaccharides such as agarose, laminarin, lichenan, and xyloglucan. In contrast, iota-carrageenases constitute a novel glycoside hydrolase structural family. We report here the crystal structure of Alteromonas fortis iota-carrageenase at 1.6 A resolution. The enzyme folds into a right-handed parallel beta-helix of 10 complete turns with two additional C-terminal domains. Glu(245), Asp(247), or Glu(310), in the cleft of the enzyme, are proposed as candidate catalytic residues. The protein contains one sodium and one chloride binding site and three calcium binding sites shown to be involved in stabilizing the enzyme structure.
- Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel, CNRS/Commissariat à l'Energie Atomique, 41, rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: