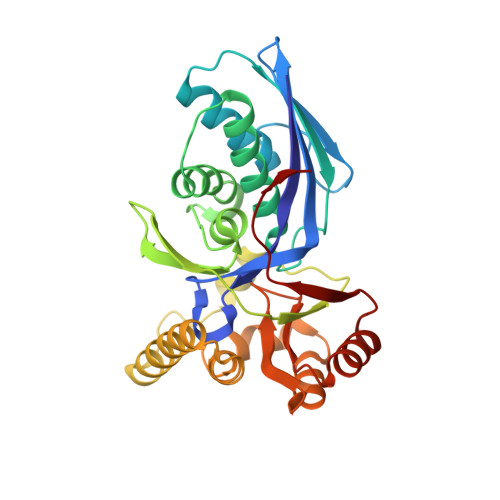
Structural Basis for the Catalysis and Substrate Specificity of Homoserine Kinase
Krishna, S.S., Zhou, T., Daugherty, M., Osterman, A.L., Zhang, H.(2001) Biochemistry 40: 10810
- PubMed: 11535056 Search on PubMed
- DOI: https://doi.org/10.1021/bi010851z
- Primary Citation Related Structures:
1H72, 1H73, 1H74 - PubMed Abstract:
Homoserine kinase (HSK), the fourth enzyme in the aspartate pathway of amino acid biosynthesis, catalyzes the phosphorylation of L-homoserine (Hse) to L-homoserine phosphate, an intermediate in the production of L-threonine, L-isoleucine, and in higher plants, L-methionine. The high-resolution structures of Methanococcus jannaschii HSK ternary complexes with its amino acid substrate and ATP analogues have been determined by X-ray crystallography. These structures reveal the structural determinants of the tight and highly specific binding of Hse, which is coupled with local conformational changes that enforce the sequestration of the substrate. The delta-hydroxyl group of bound Hse is only 3.4 A away from the gamma-phosphate of the bound nucleotide, poised for the in-line attack at the gamma-phosphorus. The bound nucleotides are flexible at the triphosphate tail. Nevertheless, a Mg(2+) was located in one of the complexes that binds between the beta- and gamma-phosphates of the nucleotide with good ligand geometry and is coordinated by the side chain of Glu130. No strong nucleophile (base) can be located near the phosphoryl acceptor hydroxyl group. Therefore, we propose that the catalytic mechanism of HSK does not involve a catalytic base for activating the phosphoryl acceptor hydroxyl but instead is mediated via a transition state stabilization mechanism.
- Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, Texas 75390-9038, USA.
Organizational Affiliation: