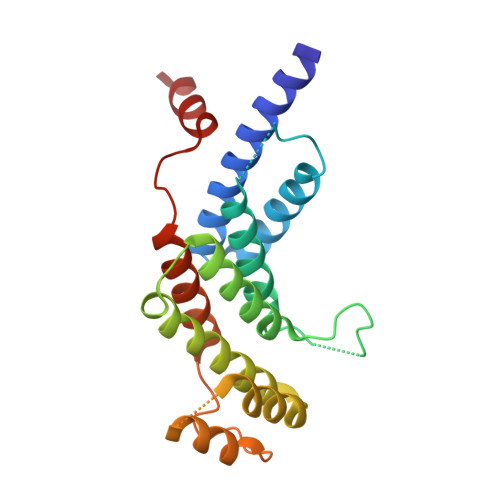
Structure of the Trfh Dimerization Domain of the Human Telomere Proteins Trf1 and Trf2
Fairall, L., Chapman, L., Moss, H., de Lange, T., Rhodes, D.(2001) Mol Cell 8: 351-361
- PubMed: 11545737
- DOI: https://doi.org/10.1016/s1097-2765(01)00321-5
- Primary Citation Related Structures:
1H6O, 1H6P - PubMed Abstract:
TRF1 and TRF2 are key components of vertebrate telomeres. They bind to double-stranded telomeric DNA as homodimers. Dimerization involves the TRF homology (TRFH) domain, which also mediates interactions with other telomeric proteins. The crystal structures of the dimerization domains from human TRF1 and TRF2 were determined at 2.9 and 2.2 A resolution, respectively. Despite a modest sequence identity, the two TRFH domains have the same entirely alpha-helical architecture, resembling a twisted horseshoe. The dimerization interfaces feature unique interactions that prevent heterodimerization. Mutational analysis of TRF1 corroborates the structural data and underscores the importance of the TRFH domain in dimerization, DNA binding, and telomere localization. A possible structural homology between the TRFH domain of fission yeast telomeric protein Taz1 with those of the vertebrate TRFs is suggested.
- MRC Laboratory of Molecular Biology, Hills Road, CB2 2QH, Cambridge, United Kingdom.
Organizational Affiliation: