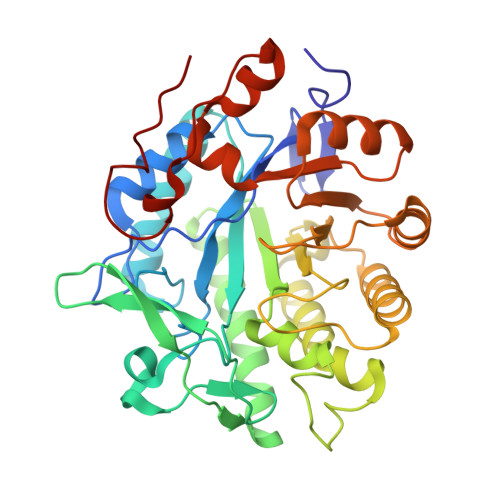
Crystal Structure of Pentaerythritol Tetranitrate Reductase: "Flipped" Binding Geometries for Steroid Substrates in Different Redox States of the Enzyme
Barna, T.M., Khan, H., Bruce, N.C., Barsukov, I., Scrutton, N.S., Moody, P.C.(2001) J Mol Biology 310: 433
- PubMed: 11428899 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4779
- Primary Citation Related Structures:
1H50, 1H51, 1H60, 1H61, 1H62, 1H63 - PubMed Abstract:
Pentaerythritol tetranitrate reductase (PETN reductase) degrades high explosive molecules including nitrate esters, nitroaromatics and cyclic triazine compounds. The enzyme also binds a variety of cyclic enones, including steroids; some steroids act as substrates whilst others are inhibitors. Understanding the basis of reactivity with cyclic enones requires structural information for the enzyme and key complexes formed with steroid substrates and inhibitors. The crystal structure of oxidised and reduced PETN reductase at 1.5 A resolution establishes a close structural similarity to the beta/alpha-barrel flavoenzyme, old yellow enzyme. In complexes of oxidised PETN reductase with progesterone (an inhibitor), 1,4-androstadiene-3,17-dione and prednisone (both substrates) the steroids are stacked over the si-face of the flavin in an orientation different from that reported for old yellow enzyme. The specifically reducible 1,2 unsaturated bonds in 1,4-androstadiene-3,17-dione and prednisone are not optimally aligned with the flavin N5 in oxidised enzyme complexes. These structures suggest either relative "flipping" or shifting of the steroid with respect to the flavin when bound in different redox forms of the enzyme. Deuterium transfer from nicotinamide coenzyme to 1,4-androstadiene-3,17-dione via the enzyme bound FMN indicates 1alpha addition at the steroid C2 atom. These studies rule out lateral motion of the steroid and indicate that the steroid orientation is "flipped" in different redox states of the enzyme.
- Department of Biochemistry, University of Leicester, Leicester, LE1 7RH, UK.
Organizational Affiliation: