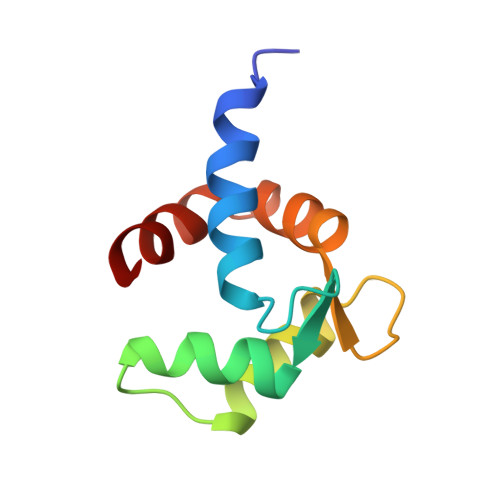
Solution Structure, Dynamics, and Hydrodynamics of the Calcium-Bound Cross-Reactive Birch Pollen Allergen Bet V 4 Reveal a Canonical Monomeric Two EF-Hand Assembly with a Regulatory Function
Neudecker, P., Nerkamp, J., Eisenmann, A., Nourse, A., Lauber, T., Schweimer, K., Lehmann, K., Schwarzinger, S., Ferreira, F., Roesch, P.(2004) J Mol Biology 336: 1141
- PubMed: 15037075
- DOI: https://doi.org/10.1016/j.jmb.2003.12.070
- Primary Citation of Related Structures:
1H4B - PubMed Abstract:
Birch pollinosis is one of the prevailing allergic diseases. In all, 5-20% of birch pollinotics mount IgE antibodies against the minor birch pollen allergen Bet v 4, a Ca2+-binding polcalcin. Due to IgE cross-reactivity among the polcalcins these patients are polysensitized to various plant pollens. Determination of the high-resolution structure of holo Bet v 4 by heteronuclear NMR spectroscopy reveals a canonical two EF-hand assembly in the open conformation with interhelical angles closely resembling holo calmodulin. The polcalcin-specific amphipathic COOH-terminal alpha-helix covers only a part of the hydrophobic groove on the molecular surface. Unlike the polcalcin Phl p 7 from timothy grass, which was recently shown to form a domain-swapped dimer, the hydrodynamic parameters from NMR relaxation, NMR translational diffusion, and analytical ultracentrifugation indicate that both apo and holo Bet v 4 are predominantly monomeric, raising the question of the physiological and immunological significance of the dimeric form of these polcalcins, whose physiological function is still unknown. The reduced helicity and heat stability in the CD spectra, the poor chemical shift dispersion of the NMR spectra, and the slightly increased hydrodynamic radius of apo Bet v 4 indicate a reversible structural transition upon Ca2+ binding, which explains the reduced IgE binding capacity of apo Bet v 4. The remarkable structural similarity of holo Bet v 4 and holo Phl p 7 in spite of different oligomerization states explains the IgE cross-reactivity and indicates that canonical monomers and domain-swapped dimers may be of similar allergenicity. Together with the close structural homology to calmodulin and the hydrophobic ligand binding groove this transition suggests a regulatory function for Bet v 4.
- Lehrstuhl für Biopolymere, Universität Bayreuth, D-95440 Bayreuth, Germany. philipp.neudecker@uni-bayreuth.de
Organizational Affiliation: