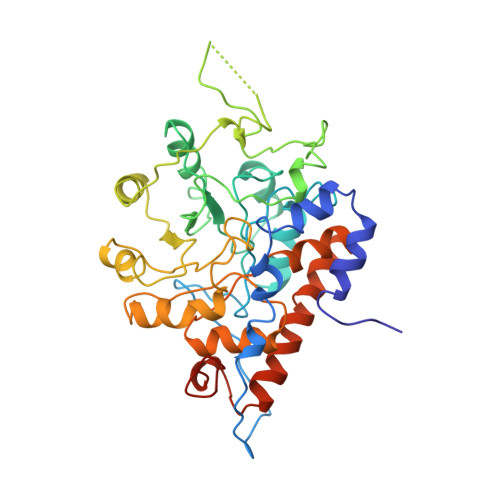
Involvement of Tyr24 and Trp108 in substrate binding and substrate specificity of glycolate oxidase.
Stenberg, K., Clausen, T., Lindqvist, Y., Macheroux, P.(1995) Eur J Biochem 228: 408-416
- PubMed: 7705356
- Primary Citation of Related Structures:
1GYL - PubMed Abstract:
Tyr24 and Trp108 are located in the active site of spinach glycolate oxidase. To elucidate their function in substrate binding and catalysis, they were replaced by phenylalanine and serine, respectively. The [Y24F]glycolate oxidase mutant enzyme showed a tenfold higher Km value for glycolate. L-lactate and DL-2-hydroxybutyrate also showed higher Km values, however, the substrate specificity was unchanged as compared to the wild-type enzyme (Km increases in the order glycolate < DL-2-hydroxybutyrate < L-lactate < L-mandelate). The turnover number and the rate of reduction, found to be rate limiting in catalysis, were only slightly affected by the deletion of the hydroxyl group. These findings suggest that Tyr24 is mostly involved in substrate binding. The spectral features of the [Y24F]glycolate oxidase suggest that a fraction (50-80%) of the protein bears a flavin N(5) adduct instead of the oxidized cofactor. Crystals obtained from the isolated [Y24F]glycolate oxidase mutant protein allowed the determination of the three-dimensional structure. Although the structure was low resolution (0.3 nm), it is evident that the structure determined is that of the N(5) adduct species. In addition to the lacking hydroxyl group of Tyr24, we also observed movements of the amino acid side chains of Arg164 and Trp108, the latter replacing a water molecule in the substrate-binding pocket. Other features predominantly found in the class of flavoprotein oxidases, such as stabilization of the covalent N(5)-sulfite adduct and of the paraquinoid form of 8-mercapto-FMN, were found to be conserved. [W108S]Glycolate oxidase, in contrast, showed dramatic effects on both the Km of substrates as well as the turnover number. The Km for glycolate was increased some hundred fold and the turnover number was decreased 500-fold. In addition, it was found that the higher homologs of glycolate, L-lactate and DL-2-hydroxybutyrate had turnover numbers similar to those found with the wild-type enzyme, although the Km values also increased dramatically. These results indicate that Trp108 is of major importance in catalysis and that this residue is involved in determining the substrate specificity of glycolate oxidase.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala Biomedical Center.
Organizational Affiliation: