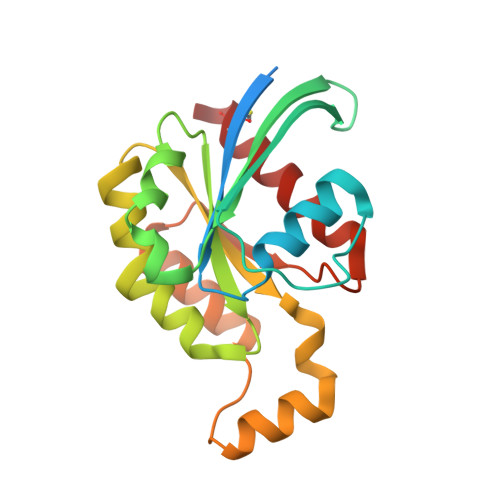
Crystal Structure of the Core Domain of Rhoe/Rnd3: A Constitutively Activated Small G Protein
Garavini, H., Riento, K., Phelan, J.P., McAlister, M.S.B., Ridley, A.J., Keep, N.H.(2002) Biochemistry 41: 6303-6310
- PubMed: 12009891
- DOI: https://doi.org/10.1021/bi025651h
- Primary Citation Related Structures:
1GWN - PubMed Abstract:
We report the 2.1 A crystal structure of the core G protein domain of the unusual Rho family member RhoE/Rnd3 in complex with endogenous GTP and magnesium. Unlike other small G proteins, RhoE, along with two other proteins Rnd1/Rho6 and Rnd2/RhoN, does not hydrolyze GTP. The main reason for this is the presence of serines in the positions equivalent to Ala59 and Gln61 in Ras. The structure shows that there are still water molecules in similar positions to the waters thought to be involved in the hydrolysis reaction in other G proteins. The structure suggests three not necessarily exclusive explanations for the lack of hydrolysis. The lack of the conserved glutamine raises the energy of the transition state inhibiting hydrolysis. The serines may restrain the waters from moving closer to the GTP, a step that is required to attain the transition state. They also stabilize the GTP-bound conformation of switch II and could prevent conformational changes required during hydrolysis. By superposition of the RhoE structure on structures of Rho family proteins in complex with binding partners, we make predictions on RhoE interactions with these partners.
- BBSRC Bloomsbury Centre for Structural Biology and School of Crystallography, Birkbeck College, University of London, Malet Street, London WC1E 7HX.
Organizational Affiliation: