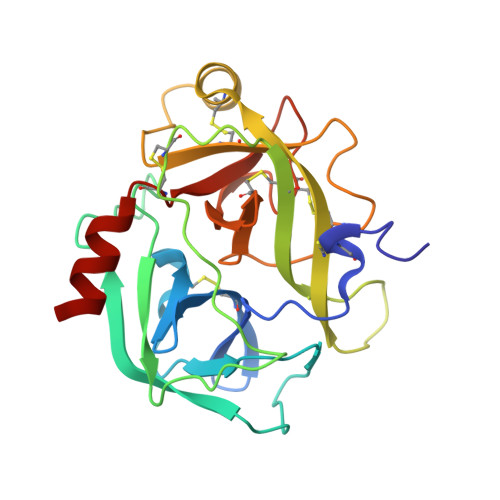
The Structure of Human Prokallikrein 6 Reveals a Novel Activation Mechanism for the Kallikrein Family.
Gomis-Ruth, F.X., Bayes, A., Sotiropoulou, G., Pampalakis, G., Tsetsenis, T., Villegas, V., Aviles, F.X., Coll, M.(2002) J Biological Chem 277: 27273
- PubMed: 12016211 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M201534200
- Primary Citation Related Structures:
1GVL - PubMed Abstract:
Zyme/protease M/neurosin/human kallikrein 6 (hK6) is a member of the human kallikrein family of trypsin-like serine proteinases and was originally identified as being down-regulated in metastatic breast and ovarian tumors when compared with corresponding primary tumors. Recent evidence suggests that hK6 may serve as a circulating tumor marker in ovarian cancers. In addition, it was described in the brain of Parkinson's disease and Alzheimer's disease patients, where it is implicated in amyloid precursor protein processing. It is thus a biomarker for these diseases. To examine the mechanism of activation of hK6, we have solved the structure of its proform, the first of a human kallikrein family member. The proenzyme displays a fold that exhibits chimeric features between those of trypsinogen and other family members. It lacks the characteristic "kallikrein loop" and forms the six disulfide bridges of trypsin. Pro-hK6 displays a completely closed specificity pocket and a unique conformation of the regions involved in structural rearrangements upon proteolytic cleavage activation. This points to a novel activation mechanism, which could be extrapolated to other human kallikreins.
- Institut de Biologia Molecular de Barcelona, Consejo Superior de Investigaciones Cientificas, c/Jordi Girona 18-26, Barcelona 08034, Spain. xgrcri@ibmb.csic.es
Organizational Affiliation: