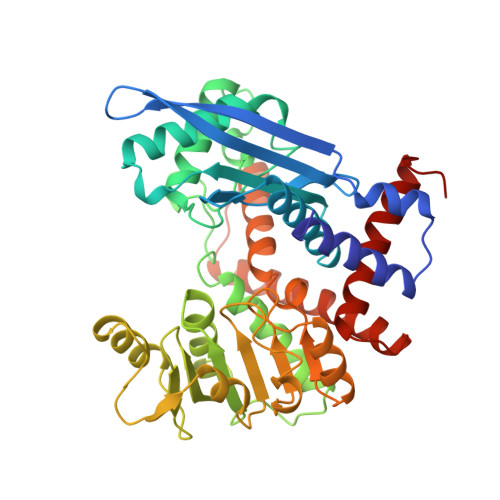
The structure of Pyrococcus furiosus glutamate dehydrogenase reveals a key role for ion-pair networks in maintaining enzyme stability at extreme temperatures.
Yip, K.S., Stillman, T.J., Britton, K.L., Artymiuk, P.J., Baker, P.J., Sedelnikova, S.E., Engel, P.C., Pasquo, A., Chiaraluce, R., Consalvi, V., Scandurra, R., Rice, D.W.(1995) Structure 3: 1147-1158
- PubMed: 8591026 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00251-9
- Primary Citation Related Structures:
1GTM, 1HRD - PubMed Abstract:
The hyperthermophile Pyrococcus furiosus is one of the most thermostable organisms known, with an optimum growth temperature of 100 degrees C. The proteins from this organism display extreme thermostability. We have undertaken the structure determination of glutamate dehydrogenase from P. furiosus in order to gain further insights into the relationship between molecular structure and thermal stability. The structure of P. furiosus glutamate dehydrogenase, a homohexameric enzyme, has been determined at 2.2 A resolution and compared with the structure of glutamate dehydrogenase from the mesophile Clostridium symbiosum. Comparison of the structures of these two enzymes has revealed one major difference: the structure of the hyperthermophilic enzyme contains a striking series of ion-pair networks on the surface of the protein subunits and buried at both interdomain and intersubunit interfaces. We propose that the formation of such extended networks may represent a major stabilizing feature associated with the adaptation of enzymes to extreme temperatures.
- The Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, University of Sheffield, PO Box 594, Sheffield S10 2UH, UK.
Organizational Affiliation: