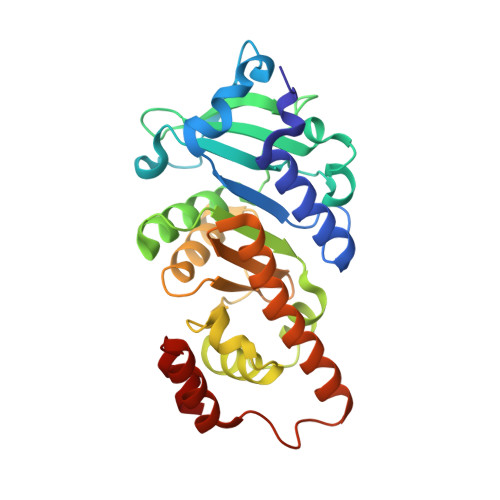
The Structure of L-Rhamnulose-1-Phosphate Aldolase (Class II) Solved by Low-Resolution Sir Phasing and 20-Fold Ncs Averaging
Kroemer, M., Schulz, G.E.(2002) Acta Crystallogr D Biol Crystallogr 58: 824
- PubMed: 11976494
- DOI: https://doi.org/10.1107/s0907444902004614
- Primary Citation of Related Structures:
1GT7 - PubMed Abstract:
The enzyme L-rhamnulose-1-phosphate aldolase catalyzes the reversible cleavage of L-rhamnulose-1-phosphate to dihydroxyacetone phosphate and L-lactaldehyde. It is a homotetramer with an M(r) of 30 000 per subunit and crystallized in space group P3(2)21. The enzyme shows a low sequence identity of 18% with the structurally known L-fuculose-1-phosphate aldolase that splits a stereoisomer in a similar reaction. Structure analysis was initiated with a single heavy-atom derivative measured to 6 A resolution. The resulting poor electron density, a self-rotation function and the working hypothesis that both enzymes are C(4) symmetric with envelopes that resemble one another allowed the location of the 20 protomers of the asymmetric unit. The crystal-packing unit was a D(4)-symmetric propeller consisting of five D(4)-symmetric octamers around an internal crystallographic twofold axis. Presumably, the propellers associate laterally in layers, which in turn pile up along the 3(2) axis to form the crystal. The non-crystallographic symmetry was used to extend the phases to the 2.7 A resolution limit and to establish a refined atomic model of the enzyme. The structure showed that the two enzymes are indeed homologous and that they possess chemically similar active centres.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität Freiburg, Albertstrasse 21, 79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: