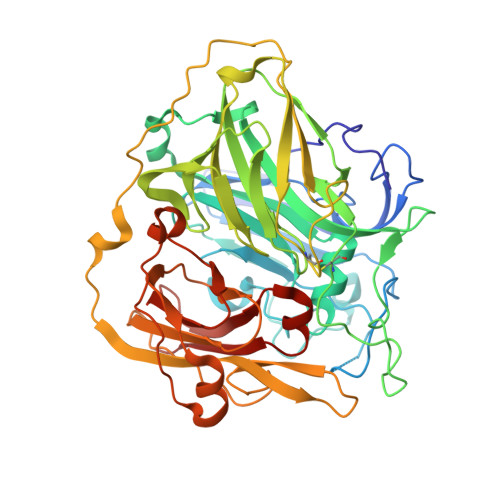
Crystal Structure of a Bacterial Endospore Coat Component: A Laccase with Enhanced Thermostability Properties
Enguita, F.J., Martins, L.O., Henriques, A.O., Carrondo, M.A.(2003) J Biological Chem 278: 19416
- PubMed: 12637519 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M301251200
- Primary Citation Related Structures:
1GSK - PubMed Abstract:
Endospores produced by the Gram-positive soil bacterium Bacillus subtilis are shielded by a proteinaceous coat formed by over 30 structural components, which self-assemble into a lamellar inner coat and a thicker striated electrodense outer coat. The 65-kDa CotA protein is an abundant component of the outer coat layer. CotA is a highly thermostable laccase, assembly of which into the coat is required for spore resistance against hydrogen peroxide and UV light. Here, we report the structure of CotA at 1.7-A resolution, as determined by x-ray crystallography. This is the first structure of an endospore coat component, and also the first structure of a bacterial laccase. The overall fold of CotA comprises three cupredoxin-like domains and includes one mononuclear and one trinuclear copper center. This arrangement is similar to that of other multicopper oxidases and most similar to that of the copper tolerance protein CueO of Escherichia coli. However, the three cupredoxin domains in CotA are further linked by external interdomain loops, which increase the packing level of the structure. We propose that these interdomain loops contribute to the remarkable thermostability of the enzyme, but our results suggest that additional factors are likely to play a role. Comparisons with the structure of other monomeric multicopper oxidases containing four copper atoms suggest that CotA may accept the largest substrates of any known laccase. Moreover, and unlike other laccases, CotA appears to have a flexible lidlike region close to the substrate-binding site that may mediate substrate accessibility. The implications of these findings for the properties of CotA, its assembly and the properties of the bacterial spore coat structure are discussed.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, 2781-901 Oeiras.
Organizational Affiliation: