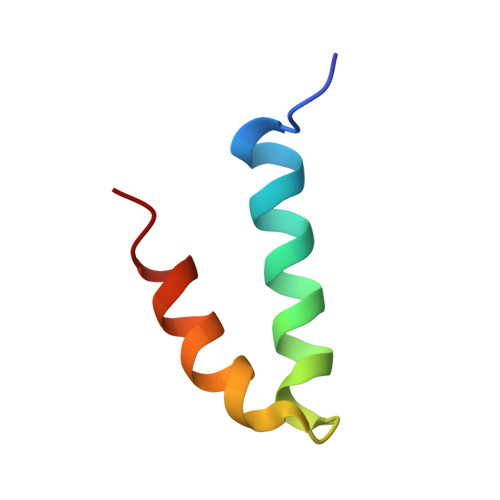
Structure of the coat protein-binding domain of the scaffolding protein from a double-stranded DNA virus.
Sun, Y., Parker, M.H., Weigele, P., Casjens, S., Prevelige Jr., P.E., Krishna, N.R.(2000) J Mol Biology 297: 1195-1202
- PubMed: 10764583
- DOI: https://doi.org/10.1006/jmbi.2000.3620
- Primary Citation Related Structures:
1GP8, 2GP8 - PubMed Abstract:
Scaffolding proteins are required for high fidelity assembly of most high T number dsDNA viruses such as the large bacteriophages, and the herpesvirus family. They function by transiently binding and positioning the coat protein subunits during capsid assembly. In both bacteriophage P22 and the herpesviruses the extreme scaffold C terminus is highly charged, is predicted to be an amphipathic alpha-helix, and is sufficient to bind the coat protein, suggesting a common mode of action. NMR studies show that the coat protein-binding domain of P22 scaffolding protein exhibits a helix-loop-helix motif stabilized by a hydrophobic core. One face of the motif is characterized by a high density of positive charges that could interact with the coat protein through electrostatic interactions. Results from previous studies with a truncation fragment and the observed salt sensitivity of the assembly process are explained by the NMR structure.
- Comprehensive Cancer Center, Birmingham, AL, 35294, USA.
Organizational Affiliation: