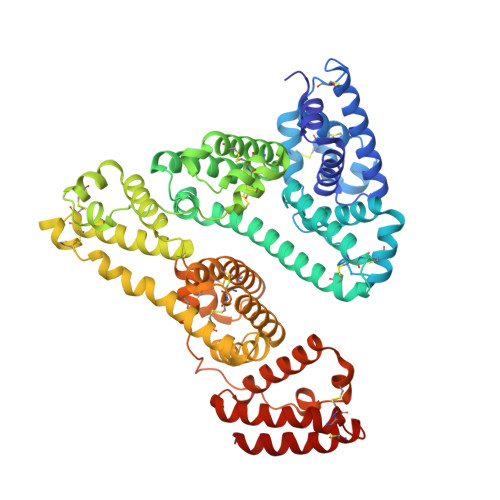
Crystal Structures of Human Serum Albumin Complexed with Monounsaturated and Polyunsaturated Fatty Acids.
Petitpas, I., Gruene, T., Bhattacharya, A.A., Curry, S.(2001) J Mol Biology 314: 955
- PubMed: 11743713 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.5208
- Primary Citation Related Structures:
1GNI, 1GNJ - PubMed Abstract:
The primary ligands of human serum albumin (HSA), an abundant plasma protein, are non-esterified fatty acids. In vivo, the majority of fatty acids associated with the protein are unsaturated. We present here the first high-resolution crystal structures of HSA complexed with two important unsaturated fatty acids, the monounsaturated oleic acid (C18:1) and the polyunsaturated arachidonic acid (C20:4). Both compounds are observed to occupy the seven binding sites distributed across the protein that are also bound by medium and long-chain saturated fatty acids. Although C18:1 fatty acid binds each site on HSA in a conformation almost identical with that of the corresponding saturated compound (C18:0), the presence of multiple cis double bonds in C20:4 induces distinct binding configurations at some sites. The observed restriction on binding configurations plausibly accounts for differences in the pattern of binding affinities for the primary sites between polyunsaturated fatty acids and their saturated or monounsaturated counterparts.
- Biophysics Section, Department of Biological Sciences, Blackett Laboratory, Imperial College of Science, Technology and Medicine, London SW7 2BW, UK.
Organizational Affiliation: