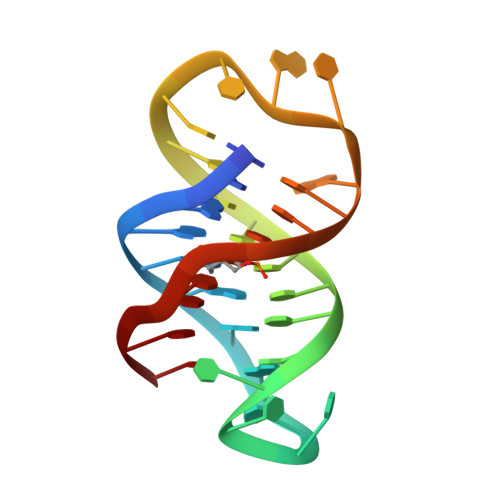
Solution structure of an intramolecular DNA triplex containing an N7-glycosylated guanine which mimics a protonated cytosine.
Koshlap, K.M., Schultze, P., Brunar, H., Dervan, P.B., Feigon, J.(1997) Biochemistry 36: 2659-2668
- PubMed: 9054573
- DOI: https://doi.org/10.1021/bi962438a
- Primary Citation Related Structures:
1GN7 - PubMed Abstract:
The three-dimensional structure of a pyrimidine-purine-pyrimidine DNA triplex containing an N7-glycosylated guanine (7G) in the third strand has been determined by NMR spectroscopy, restrained molecular dynamics calculations, and complete relaxation matrix refinement. Glycosylation of the guanine at the N7 position permits it to adopt a conformation such that the Hoogsteen face of the base mimics the arrangement of hydrogen bond donors seen in protonated cytosine. The NMR data confirm the previously proposed hydrogen bonding scheme of the 7G x G x C triplet. The three-dimensional structure of the triplex accommodates the 7G with less distortion of the phosphodiester backbone than would be required for an N9-glycosylated guanine in the same sequence position, but some changes in the positions of the phosphodiester backbone are present compared to a C+ x G x C triplet. The structure provides a rationale for the observations that 7G binds to Watson-Crick G x C base pairs with higher specificity and affinity than guanine, but with a lower stability at pH 5.2 than would be provided by a canonical C+ x G x C triplet.
- Department of Chemistry and Biochemistry, University of California, Los Angeles 90095, USA.
Organizational Affiliation: