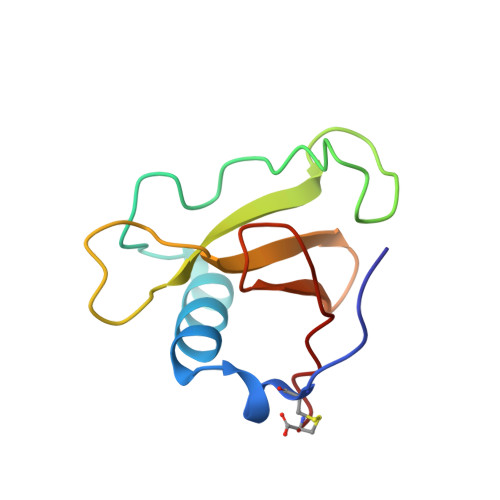
Complex of ribonuclease from Streptomyces aureofaciens with 2'-GMP at 1.7 A resolution.
Sevcik, J., Hill, C.P., Dauter, Z., Wilson, K.S.(1993) Acta Crystallogr D Biol Crystallogr 49: 257-271
- PubMed: 15299531 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444992007261
- Primary Citation Related Structures:
1GMP, 1GMQ, 1GMR - PubMed Abstract:
The crystal structure of a complex of ribonuclease from Streptomyces aureofaciens (RNase Sa) with guanosine-2'-monophosphate (2'-GMP) has been refined against synchrotron data recorded from a single crystal using radiation from beamline X31 at EMBL, Hamburg, and an imaging plate scanner. The crystals are in space group P2(1)2(1)2(1) with cell dimensions a = 64.7, b = 78.8 and c = 39.1 A. The structure has two enzyme molecules in the asymmetric unit, complexed with 2'-GMP inhibitor with occupancies of 1 and 2/3 (different to the 3'-GMP complex crystal structure where only one of the two independent RNase Sa molecules binds nucleotide), 492 associated water molecules and one sulfate ion, and was refined using all data between 10.0 and 1.7 A to a final crystallographic R factor of 13.25%. Binding of the base to the enzyme confirms the basis for the guanine specificity but the structural results still do not provide direct evidence of the identity and role of the particular residues involved in the catalytic process. New native RNase Sa data to 1.8 A were recorded to provide a reference set measured under comparable experimental conditions. The crystals are in the same space group and have the same lattice as those of the 2'-GMP complex. The native structure with 423 water molecules was refined in a similar manner to the complex to a final R factor of 13.87%. 1.77 A resolution data were independently measured on a 2'-GMP complex crystal at UCLA using an R-AXIS II image plate scanner mounted on a conventional source. The cell dimensions were essentially the same as above. 2'-GMP was bound more fully to molecule A than to molecule B of the RNase Sa. The structure was refined to an R factor of 14.64% with 388 water molecules. This work follows on from the structure determination of native RNase Sa and its complex with 3'-GMP [Sevcik, Dodson & Dodson (1991). Acta Cryst. B47, 240-253].
- Institute of Molecular Biology, Slovak Academy of Sciences, Bratislava, Czechoslovakia.
Organizational Affiliation: