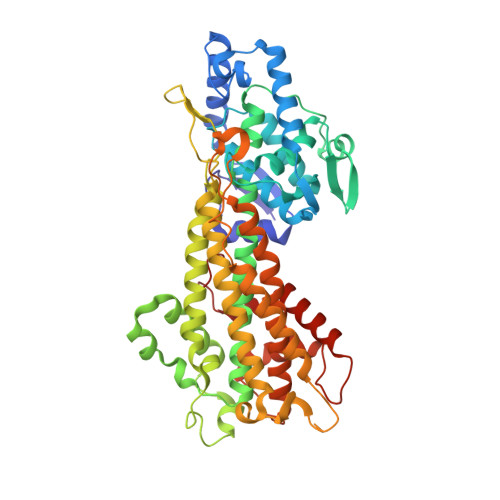
Autocatalytic Peptide Cyclization During Chain Folding of Histidine Ammonia-Lyase.
Baedeker, M., Schulz, G.E.(2002) Structure 10: 61-67
- PubMed: 11796111 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00692-x
- Primary Citation Related Structures:
1EB4, 1GK2, 1GK3 - PubMed Abstract:
Histidine ammonia-lyase requires a 4-methylidene-imidazole-5-one group (MIO) that is produced autocatalytically by a cyclization and dehydration step in a 3-residue loop of the polypeptide. The crystal structures of three mutants have been established. Two mutants were inactive and failed to form MIO, but remained unchanged elsewhere. The third mutant showed very low activity and formed MIO, although it differed from an MIO-less mutant only by an additional 329-C(beta) atom. This atom forms one constraint during MIO formation, the other being the strongly connected Asp145. An exploration of the conformational space of the MIO-forming loop showed that the cyclization is probably enforced by a mechanic compression in a late stage of chain folding and is catalyzed by a well-connected internal water molecule. The cyclization of the respective 3-residue loop of green fluorescent protein is likely to occur in a similar reaction.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, D-79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: