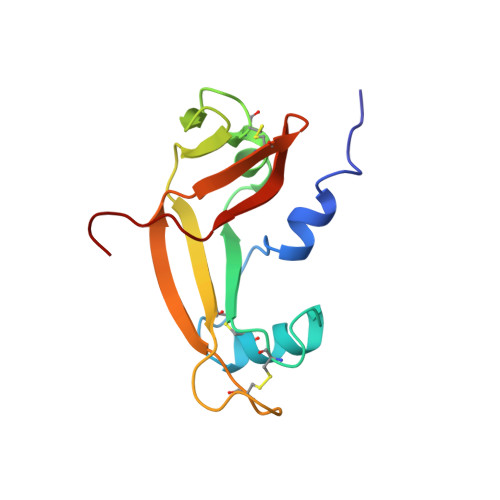
Solution structure of bovine angiogenin by 1H nuclear magnetic resonance spectroscopy.
Lequin, O., Albaret, C., Bontems, F., Spik, G., Lallemand, J.Y.(1996) Biochemistry 35: 8870-8880
- PubMed: 8688423 Search on PubMed
- DOI: https://doi.org/10.1021/bi960022r
- Primary Citation Related Structures:
1GIO - PubMed Abstract:
The three-dimensional structure of bovine angiogenin has been determined using two- and three-dimensional proton NMR spectroscopy. The solution structure is very close to that recently determined by X-ray diffraction analysis. This structure appears well defined, even if five loops and one helix exhibit greater flexibility. Analysis of the active site geometry confirms the position of the Glu-118 residue which obstructs the pyrimidine binding site. There is no experimental evidence of an unobstructed conformation of angiogenin in solution. In addition, it appears that the Glu-118 and Ser-119 residues and the cell receptor binding loop may play an important role in the differences of C-terminal fragment organization and ribonucleolytic activity observed between angiogenins and ribonuclease A.
- Départment de Chimie-Synthèse Organique, URA 1308 du CNRS, Ecole Polytechnique, Palaiseau, France.
Organizational Affiliation: