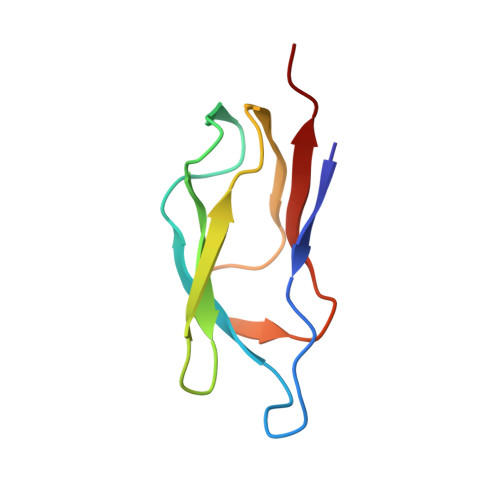
Solution structure of the lipoyl domain of the 2-oxoglutarate dehydrogenase complex from Azotobacter vinelandii.
Berg, A., Vervoort, J., de Kok, A.(1996) J Mol Biology 261: 432-442
- PubMed: 8780784 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0474
- Primary Citation Related Structures:
1GHJ, 1GHK - PubMed Abstract:
The three-dimensional solution structure of the lipoyl domain of the 2-oxoglutarate dehydrogenase complex from Azotobacter vinelandii has been determined from nuclear magnetic resonance data by using distance geometry and dynamical simulated annealing refinement. The structure determination is based on a total of 580 experimentally derived distance constraints and 65 dihedral angle constraints. The solution structure is represented by an ensemble of 25 structures with an average root-mean-square deviation between the individual structures of the ensemble and the mean coordinates of 0.71 A for backbone atoms and 1.08 A for all heavy atoms. The overall fold of the lipoyl domain is that of a beta-barrel-sandwich hybrid. It consists of two almost parallel four-stranded anti-parallel beta-sheets formed around a well-defined hydrophobic core, with a central position of the single tryptophan 21. The lipoylation site, lysine 42, is found in a beta-turn at the far end of one of the sheets, and is close in space to a solvent-exposed loop comprising residues 7 to 15. The lipoyl domain displays a remarkable internal symmetry that projects one beta-sheet onto the other beta-sheet after rotation of approximately 180 degrees about a 2-fold rotational symmetry axis. There is close structural similarity between the structure of this 2-oxoglutarate dehydrogenase complex lipoyl domain and the structures of the lipoyl domains of pyruvate dehydrogenase complexes from Bacillus stearothermophilus and Escherichia coli, and conformational differences occur primarily in a solvent-exposed loop close in space to the lipoylation site. The lipoyl domain structure is discussed in relation to the process of molecular recognition of lipoyl domains by their parent 2-oxo acid dehydrogenase.
- Department of Biochemistry Agricultural University, Wageningen, The Netherlands.
Organizational Affiliation: