A STRUCTURAL EXPLANATION FOR ENZYME MEMORY IN NONAQUEOUS SOLVENTS.
Yennawar, H.P., Yennawar, N.H., Farber, G.K.(1995) J Am Chem Soc 117: 577-585
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(1995) J Am Chem Soc 117: 577-585
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| GAMMA-CHYMOTRYPSIN A | A [auth E] | 13 | Bos taurus | Mutation(s): 0 EC: 3.4.21.1 |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00766 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| GAMMA-CHYMOTRYPSIN A | B [auth F] | 131 | Bos taurus | Mutation(s): 0 EC: 3.4.21.1 | 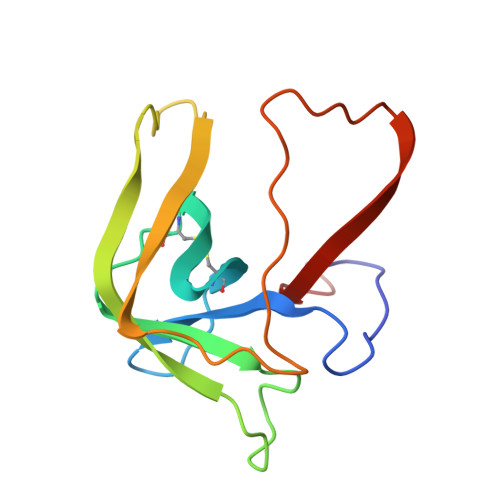 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00766 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| GAMMA-CHYMOTRYPSIN A | C [auth G] | 97 | Bos taurus | Mutation(s): 0 EC: 3.4.21.1 | 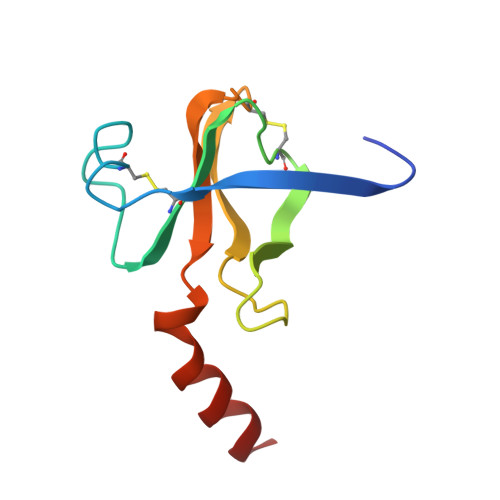 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00766 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| PRO GLY VAL TYR PEPTIDE | D [auth P] | 4 | N/A | Mutation(s): 0 | 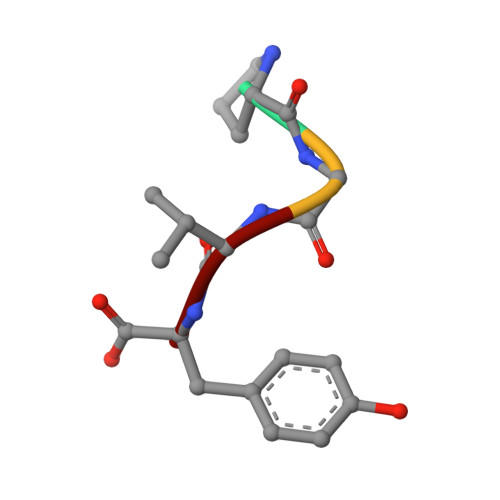 |
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Download:Ideal Coordinates CCD File | E [auth F] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| IPA Download:Ideal Coordinates CCD File | F [auth G] | ISOPROPYL ALCOHOL C3 H8 O KFZMGEQAYNKOFK-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 69.6 | α = 90 |
| b = 69.6 | β = 90 |
| c = 97.51 | γ = 90 |
| Software Name | Purpose |
|---|---|
| X-PLOR | refinement |