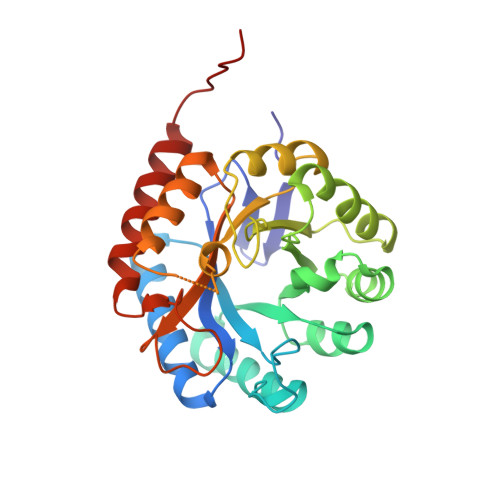
3-Deoxy-D-manno-octulosonate-8-phosphate synthase from Escherichia coli. Model of binding of phosphoenolpyruvate and D-arabinose-5-phosphate.
Wagner, T., Kretsinger, R.H., Bauerle, R., Tolbert, W.D.(2000) J Mol Biology 301: 233-238
- PubMed: 10926505 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3956
- Primary Citation Related Structures:
1GG0 - PubMed Abstract:
The crystal structure of 3-deoxy-d-manno-octulosonate-8-phosphate synthase (KDOPS) from Escherichia coli was determined by molecular replacement using coordinates given to us by Radaev and co-workers prior to publication. The KDOPS crystals reported by Radaev et al. were grown in the presence of 1.4 M (NH(4))(2)SO(4) and 0.4 M (K/H)(3)PO(4). They are in the cubic space group I23 (a=228.6 A) with a tetramer in the asymmetric unit; the structure has been refined with data to 2.4 A. Our crystals of E. coli KDOPS, grown in 24 % (w/v) polyethylene glycol (PEG) 1500 in the presence of the substrates, 2-phosphoenolpyruvate (PEP) and d-arabinose-5-phosphate (A5P), are also in space group I23 (a=118.2 A), with one subunit in the asymmetric unit. The medium of crystallization, 1.8 M SO(4)/PO(4) versus 24 % PEG, does not significantly affect the conformation of KDOPS. The inter-monomer contacts in both structures are the same. The beta(8)/alpha(8) loop (residues 246 to 251) situated near the entrance to the active site is not seen in the 229 A structure but can be traced in the 118 A structure. Most significantly, Radaev et al. interpreted two SO(4)/PO(4) sites in the 229 A structure as marking the phosphate positions of the substrates, PEP and A5P, after the precedent of DAHPS. In the 118 A structure the inner of these two SO(4)/PO(4) peaks is present at the same position as in the 229 A structure of KDOPS. The outer phosphate peak in the 118 A KDOPS is 3.7 A from the outer SO(4)/PO(4) peak in the 229 A structure and is within hydrogen bonding distance of Arg63 of the same subunit and Arg120 of another subunit. Based on the precedent of the d-erythrose-4-phosphate (E4P) modeled in the active site of DAHPS, we have modeled PEP and A5P in KDOPS and compared the coordination of PEP and A5P in KDOPS with that of PEP and E4P in DAHPS.
- Department of Biology, University of Virginia, Charlottesville, VA, 22903-2477, USA.
Organizational Affiliation: