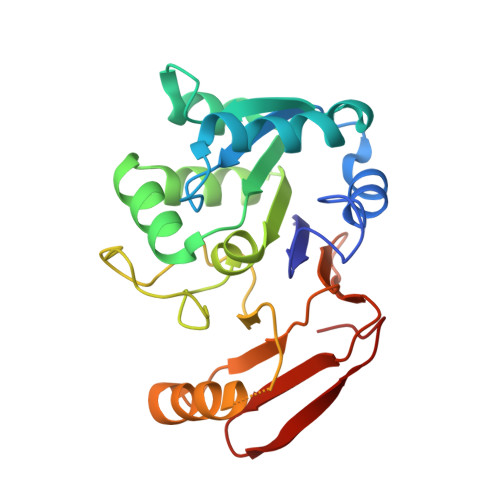
The structure of aspartyl dipeptidase reveals a unique fold with a Ser-His-Glu catalytic triad.
Hakansson, K., Wang, A.H., Miller, C.G.(2000) Proc Natl Acad Sci U S A 97: 14097-14102
- PubMed: 11106384 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.260376797
- Primary Citation Related Structures:
1FY2, 1FYE - PubMed Abstract:
The three-dimensional structure of Salmonella typhimurium aspartyl dipeptidase, peptidase E, was solved crystallographically and refined to 1.2-A resolution. The structure of this 25-kDa enzyme consists of two mixed beta-sheets forming a V, flanked by six alpha-helices. The active site contains a Ser-His-Glu catalytic triad and is the first example of a serine peptidase/protease with a glutamate in the catalytic triad. The active site Ser is located on a strand-helix motif reminiscent of that found in alpha/beta-hydrolases, but the polypeptide fold and the organization of the catalytic triad differ from those of the known serine proteases. This enzyme is a member of a family of serine hydrolases and appears to represent a new example of convergent evolution of peptidase activity.
- Departments of Microbiology and Cell and Structural Biology, B103 Chemical and Life Science Laboratory, University of Illinois at Urbana-Champaign, 601 South Goodwin Avenue, Urbana, IL 61801, USA.
Organizational Affiliation: