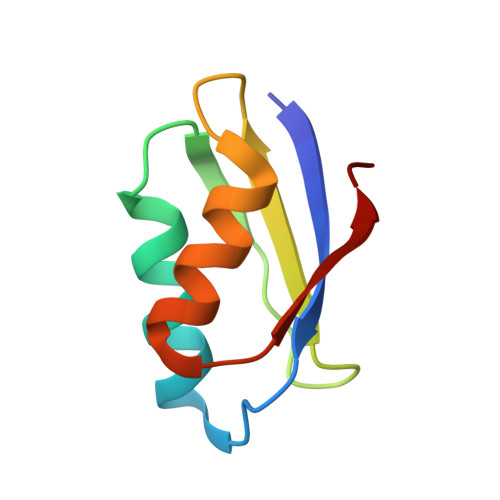
Solution structure of the yeast copper transporter domain Ccc2a in the apo and Cu(I)-loaded states.
Banci, L., Bertini, I., Ciofi-Baffoni, S., Huffman, D.L., O'Halloran, T.V.(2001) J Biological Chem 276: 8415-8426
- PubMed: 11083871 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M008389200
- Primary Citation Related Structures:
1FVQ, 1FVS - PubMed Abstract:
Ccc2 is an intracellular copper transporter in Saccharomyces cerevisiae and is a physiological target of the copper chaperone Atx1. Here we describe the solution structure of the first N-terminal MTCXXC metal-binding domain, Ccc2a, both in the presence and absence of Cu(I). For Cu(I)-Ccc2a, 1944 meaningful nuclear Overhauser effects were used to obtain a family of 35 structures with root mean square deviation to the average structure of 0.36 +/- 0.06 A for the backbone and 0.79 +/- 0.05 A for the heavy atoms. For apo-Ccc2a, 1970 meaningful nuclear Overhauser effects have been used with 35 (3)J(HNHalpha) to obtain a family of 35 structures with root mean square deviation to the average structure of 0.38 +/- 0.06 A for the backbone and 0.82 +/- 0.07 A for the heavy atoms. The protein exhibits a betaalphabetabetaalphabeta, ferrodoxin-like fold similar to that of its target Atx1 and that of a human counterpart, the fourth metal-binding domain of the Menkes protein. The overall fold remains unchanged upon copper loading, but the copper-binding site itself becomes less disordered. The helical context of the copper-binding site, and the copper-induced conformational changes in Ccc2a differ from those in Atx1. Ccc2a presents a conserved acidic surface which complements the basic surface of Atx1 and a hydrophobic surface. These results open new mechanistic aspects of copper transporter domains with physiological copper donor and acceptor proteins.
- Magnetic Resonance Center and Department of Chemistry, University of Florence, Via Luigi Sacconi 6, Sesto Fiorentino, Florence, 50019, Italy.
Organizational Affiliation: