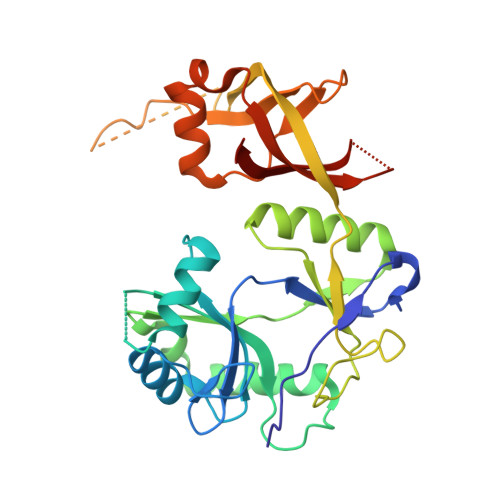
Crystal structure of eukaryotic DNA ligase-adenylate illuminates the mechanism of nick sensing and strand joining.
Odell, M., Sriskanda, V., Shuman, S., Nikolov, D.B.(2000) Mol Cell 6: 1183-1193
- PubMed: 11106756 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(00)00115-5
- Primary Citation Related Structures:
1FVI - PubMed Abstract:
Chlorella virus DNA ligase is the smallest eukaryotic ATP-dependent ligase known; it has an intrinsic nick-sensing function and suffices for yeast cell growth. Here, we report the 2.0 A crystal structure of the covalent ligase-AMP reaction intermediate. The conformation of the adenosine nucleoside and contacts between the enzyme and the ribose sugar have undergone a significant change compared to complexes of T7 ligase with ATP or mRNA capping enzyme with GTP. The conformational switch allows the 3' OH of AMP to coordinate directly the 5' PO(4) of the nick. The structure explains why nick sensing is restricted to adenylated ligase and why the 5' phosphate is required for DNA binding. We identify a metal binding site on ligase-adenylate and propose a mechanism of nick recognition and catalysis supported by mutational data.
- Molecular Biology Program and Cellular Biochemistry and Biophysics Program Sloan-Kettering Institute, New York, NY 10021, USA.
Organizational Affiliation: