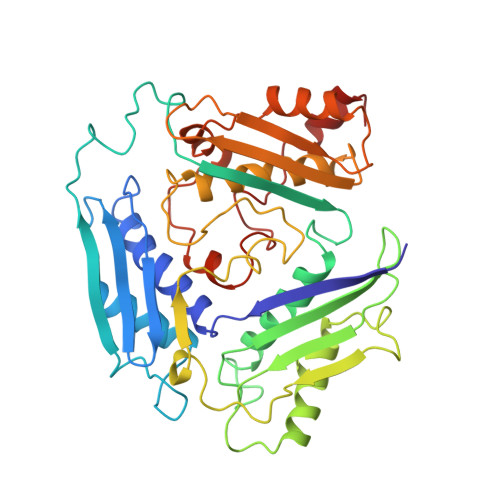
Flexible loop in the structure of S-adenosylmethionine synthetase crystallized in the tetragonal modification.
Fu, Z., Hu, Y., Markham, G.D., Takusagawa, F.(1996) J Biomol Struct Dyn 13: 727-739
- PubMed: 8723769
- DOI: https://doi.org/10.1080/07391102.1996.10508887
- Primary Citation of Related Structures:
1FUG - PubMed Abstract:
S-Adenosylmethionine synthetase (MAT, ATP:L-methionine S-adenosyltransferase, E.C.2.5.1.6.) plays a central metabolic role in all organisms. MAT catalyzes the two-step reaction which synthesizes S-adenosylmethionine (AdoMet), pyrophosphate (PPi) and orthophosphate (Pi) from ATP and L-methionine. AdoMet is the primary methyl group donor in biological systems. MAT from Escherichia coli was crystallized in the tetragonal modification with space group P4(3)2(1)2 using the same conditions as previously yielded crystals of the hexagonal system [Takusagawa, et al., (1996), J. Biol. Chem. 171, 136-147], except for the crystallization temperature. The structure has been determined by molecular replacement at 3.2 A resolution. The overall structure of the tetrameric MAT in the tetragonal modification is essentially the same as the structure found in the hexagonal modification. However there are two remarkable differences between the structures of two modifications. One is the contents in the active sites (holoform vs. apo-form), and the other is the conformation of the flexible loop over the active site (open vs. closed). These differences in the crystal structures are caused solely by the difference in crystallization temperatures (26 degrees C vs. 4 degrees C). We have interpreted the structural data obtained from the X-ray analyses in conjunction with the results of the mechanistic and sequencing studies in terms of possible dynamic motion of the flexible loop. When a substrate/product binds in the active site (hexagonal modification), the loop becomes disordered, apparently due to flexibility at the entrance of the active site as if it acts as a "mobile loop" during the catalytic reaction. On the other hand, when the temperature is decreased, the dynamic motion of the flexible loop may be reduced, and the loop residues enter the active site and close its entrance (tetragonal modification). Thus, the active site of the tetragonal modification is empty despite the crystals being grown in mother liquor containing a large concentration of phosphate (100 mM). There is no significant displacement of amino acid residues in the active site between the holo and apo forms, suggesting that the flexible loop plays an important role in determination of the contents in the active site. Since the functionally important amino acid residues in the active site are all conserved throughout various species, the structures of the active sites and the mechanism of the catalysis are probably essentially identical in the enzymes from a wide range of organisms. However, the substrate KM and Vmax values of MATs from various species are distributed over a wide range. The amino acid residues in the flexible loop regions are poorly conserved throughout various species. Therefore, the wide differences in catalysis rates of MATs from various speeches may be due to the differences in the composition of the flexible loop.
- Department of Chemistry, University of Kansas, Lawrence 66045-0046, USA.
Organizational Affiliation: