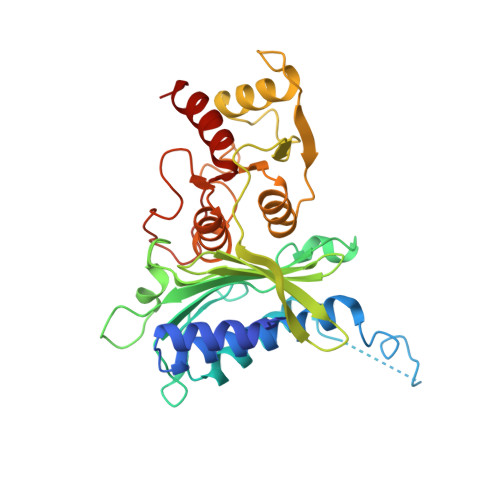
The allosteric site of human liver fructose-1,6-bisphosphatase. Analysis of six AMP site mutants based on the crystal structure.
Gidh-Jain, M., Zhang, Y., van Poelje, P.D., Liang, J.Y., Huang, S., Kim, J., Elliott, J.T., Erion, M.D., Pilkis, S.J., Raafat el-Maghrabi, M.(1994) J Biological Chem 269: 27732-27738
- PubMed: 7961695 Search on PubMed
- Primary Citation Related Structures:
1FTA - PubMed Abstract:
The molecular structure of human liver fructose-1,6-bisphosphatase complexed with AMP was determined by x-ray diffraction using molecular replacement, starting from the pig kidney enzyme AMP complex. Of the 34 amino acid residues which differ between these two sequences, only one interacts with AMP; Met30 in pig kidney is Leu30 in human liver. From this analysis, six sites in which side chains of amino acid residues are in contact with AMP, Ala24, Leu30, Thr31, Tyr113, Arg140, and Met177, were mutated by polymerase chain reaction. The wild-type and mutant forms were expressed in Escherichia coli, purified, and their kinetic properties determined. Circular dichroism spectra of the mutants were indistinguishable from that of the wild-type enzyme. Kinetic analyses revealed that all forms had similar turnover numbers, Km values for fructose 2,6-bisphosphate, and inhibition constants for fructose 2,6-bisphosphate. Apparent Ki values for AMP inhibition of the Leu30 --> Phe and Met177 --> Ala mutants were similar to those of the wild-type enzyme, but the apparent Ki values for the Arg140 --> Ala and Ala24 --> Phe mutants were 7-to 20-fold higher, respectively. The Thr31 --> Ser mutant exhibited a 5-fold increase in apparent Ki for AMP, while mutation of Thr31 to Ala increased the apparent Ki 120-fold. AMP inhibition of the Tyr113 --> Phe mutant was undetectable even at millimolar AMP concentrations. Fructose 2,6-bisphosphate potentiated AMP inhibition of the mutants to the same extent as for the wild-type enzyme, except in the case of the Thr31 --> Ala and Tyr113 --> Phe mutants. Thus, the Met177 --> Ala mutant suggests that the side chain beyond C alpha is not needed for AMP binding, and that the Leu30 --> Phe mutant preserves the AMP contacts with these side chains. Thr31, Tyr113, and Arg140 form key hydrogen bonds to AMP consistent with strong side chain interactions in the wild-type enzyme. Finally, the absence of any effect of fructose 2,6-bisphosphate on AMP inhibition observed in the Thr31 --> Ala mutant may be an important clue relating to the mechanism of synergism of these two inhibitors.
- Department of Physiology and Biophysics, State University of New York at Stony Brook 11794-8861.
Organizational Affiliation: