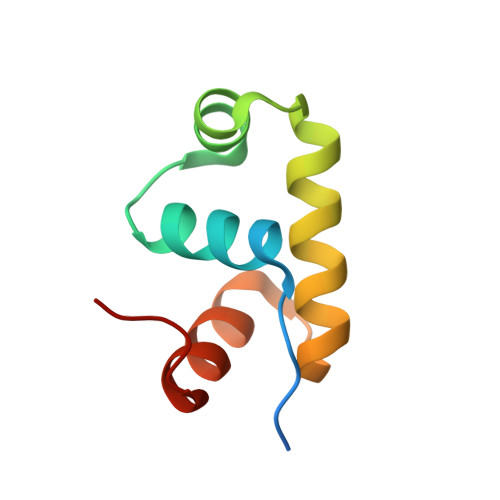
Crystal structure of GerE, the ultimate transcriptional regulator of spore formation in Bacillus subtilis.
Ducros, V.M., Lewis, R.J., Verma, C.S., Dodson, E.J., Leonard, G., Turkenburg, J.P., Murshudov, G.N., Wilkinson, A.J., Brannigan, J.A.(2001) J Mol Biology 306: 759-771
- PubMed: 11243786 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4443
- Primary Citation Related Structures:
1FSE - PubMed Abstract:
The small, DNA-binding protein GerE regulates gene transcription in the terminally differentiated mother-cell compartment during late stages of sporulation in Bacillus subtilis. This versatile transcription factor shares sequence homology with the LuxR/FixJ/UhpA family of activators and modulates the expression of a number of genes, in particular those encoding the components of the coat that surrounds the mature spore. GerE orchestrates the final stages of coat deposition and maturation that lead to a spore with remarkable resistance properties but that must be responsive to low levels of germination signals. As this germination process is largely passive and can occur in the absence of de novo protein synthesis, the correct assembly of germination machinery, including germinant receptors and energy storage compounds, is crucial to the survival of the cell. The crystal structure of GerE has been solved at 2.05 A resolution using multi-wavelength anomalous dispersion techniques and reveals the nature of the GerE dimer. Each monomer comprises four alpha-helices, of which the central pair forms a helix-turn-helix DNA-binding motif. Implications for DNA-binding and the structural organisation of the LuxR/FixJ/UhpA family of transcription activator domains are discussed.
- Structural Biology Laboratory Department of Chemistry, University of York, Heslington, YO10 5DD, UK.
Organizational Affiliation: