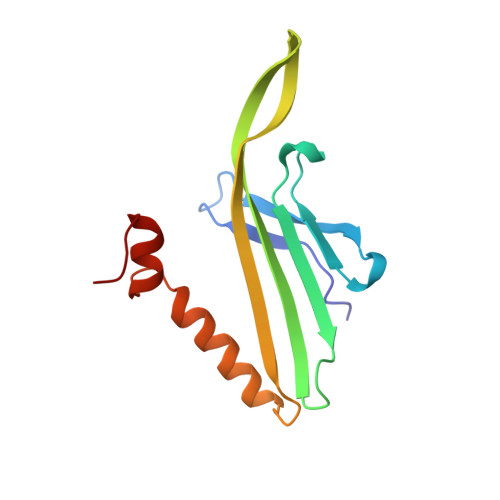
Crystal structure of bacteriophage fr capsids at 3.5 A resolution.
Liljas, L., Fridborg, K., Valegard, K., Bundule, M., Pumpens, P.(1994) J Mol Biology 244: 279-290
- PubMed: 7966339
- DOI: https://doi.org/10.1006/jmbi.1994.1729
- Primary Citation of Related Structures:
1FRS - PubMed Abstract:
The structure of recombinant capsids of the bacterial virus fr has been determined by X-ray crystallography at 3.5 A resolution. The capsids were produced by expressing the fr coat protein in Escherichia coli, the natural host of the virus, and are probably essentially identical to the protein shell of the native virus. The structure was determined using molecular replacement with the protein shell of the related MS2 virus, and refined to a crystallographic R-factor of 0.228. A comparison of the protein shells of the viruses shows that they are very similar, and indicates that they may have a similar regulation of the assembly of the quasi-symmetrical protein shell.
- Department of Molecular Biology, Uppsala University, Sweden.
Organizational Affiliation: