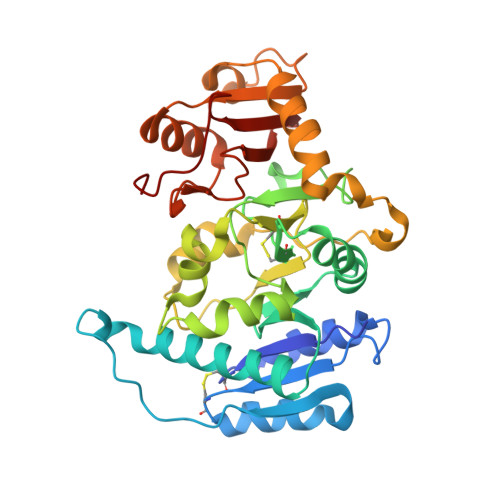
X-ray crystal structure of rabbit N-acetylglucosaminyltransferase I: catalytic mechanism and a new protein superfamily.
Unligil, U.M., Zhou, S., Yuwaraj, S., Sarkar, M., Schachter, H., Rini, J.M.(2000) EMBO J 19: 5269-5280
- PubMed: 11032794
- DOI: https://doi.org/10.1093/emboj/19.20.5269
- Primary Citation of Related Structures:
1FO8, 1FO9, 1FOA - PubMed Abstract:
N:-acetylglucosaminyltransferase I (GnT I) serves as the gateway from oligomannose to hybrid and complex N:-glycans and plays a critical role in mammalian development and possibly all metazoans. We have determined the X-ray crystal structure of the catalytic fragment of GnT I in the absence and presence of bound UDP-GlcNAc/Mn(2+) at 1.5 and 1.8 A resolution, respectively. The structures identify residues critical for substrate binding and catalysis and provide evidence for similarity, at the mechanistic level, to the deglycosylation step of retaining beta-glycosidases. The structuring of a 13 residue loop, resulting from UDP-GlcNAc/Mn(2+) binding, provides an explanation for the ordered sequential 'Bi Bi' kinetics shown by GnT I. Analysis reveals a domain shared with Bacillus subtilis glycosyltransferase SpsA, bovine beta-1,4-galactosyl transferase 1 and Escherichia coli N:-acetylglucosamine-1-phosphate uridyltransferase. The low sequence identity, conserved fold and related functional features shown by this domain define a superfamily whose members probably share a common ancestor. Sequence analysis and protein threading show that the domain is represented in proteins from several glycosyltransferase families.
- Department of Medical Genetics and Microbiology, University of Toronto, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: