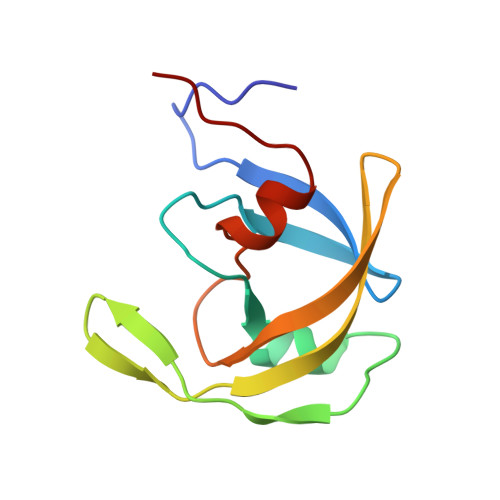
Structure of equine infectious anemia virus proteinase complexed with an inhibitor.
Gustchina, A., Kervinen, J., Powell, D.J., Zdanov, A., Kay, J., Wlodawer, A.(1996) Protein Sci 5: 1453-1465
- PubMed: 8844837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560050802
- Primary Citation Related Structures:
1FMB - PubMed Abstract:
Equine infectious anemia virus (EIAV), the causative agent of infectious anemia in horses, is a member of the lentiviral family. The virus-encoded proteinase (PR) processes viral polyproteins into functional molecules during replication and it also cleaves viral nucleocapsid protein during infection. The X-ray structure of a complex of the 154G mutant of EIAV PR with the inhibitor HBY-793 was solved at 1.8 A resolution and refined to a crystallographic R-factor of 0.136. The molecule is a dimer in which the monomers are related by a crystallographic twofold axis. Although both the enzyme and the inhibitor are symmetric, the interactions between the central part of the inhibitor and the active site aspartates are asymmetric, and the inhibitor and the two flaps are partially disordered. The overall fold of EIAV PR is very similar to that of other retroviral proteinases. However, a novel feature of the EIAV PR structure is the appearance of the second alpha-helix in the monomer in a position predicted by the structural template for the family of aspartic proteinases. The parts of the EIAV PR with the highest resemblance to human immunodeficiency virus type 1 PR include the substrate-binding sites; thus, the differences in the specificity of both enzymes have to be explained by enzyme-ligand interactions at the periphery of the active site as well.
- Macromolecular Structure Laboratory, NCI-Frederick Cancer Research and Development Center, Maryland 21702, USA.
Organizational Affiliation: