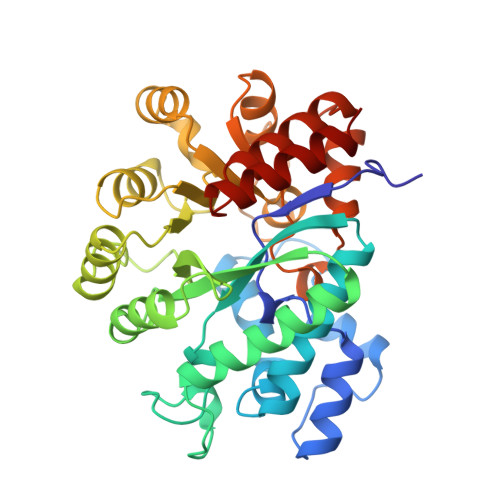
Probing the functional role of two conserved active site aspartates in mouse adenosine deaminase.
Sideraki, V., Mohamedali, K.A., Wilson, D.K., Chang, Z., Kellems, R.E., Quiocho, F.A., Rudolph, F.B.(1996) Biochemistry 35: 7862-7872
- PubMed: 8672487 Search on PubMed
- DOI: https://doi.org/10.1021/bi952920d
- Primary Citation Related Structures:
1FKW, 1FKX - PubMed Abstract:
Two adjacent aspartates, Asp 295 and Asp 296, playing major roles in the reaction catalyzed by mouse adenosine deaminase (mADA) were altered using site-directed mutagenesis. These mutants were expressed and purified from an ADA-deficient bacterial strain and characterized. Circular dichroism spectroscopy shows the mutants to have unperturbed secondary structure. Their zinc content compares well to that of wild-type enzyme. Changing Asp 295 to a glutamate decreases the kcat but does not alter the Km for adenosine, confirming the importance of this residue in the catalytic process and its minimal role in substrate binding. The crystal structure of the D295E mutant reveals a displacement of the catalytic water from the active site due to the longer glutamate side chain, resulting in the mutant's inability to turn over the substrate. In contrast, Asp 296 mutants exhibit markedly increased Km values, establishing this residue's critical role in substrate binding. The Asp 296->Ala mutation causes a 70-fold increase in the Km for adenosine and retains 0.001% of the wild-type kcat/Km value, whereas the ASP 296->Asn mutant has a 10-fold higher Km and retains 1% of the wild-type kcat/Km value. The structure of the D296A mutant shows that the impaired binding of substrate is caused by the loss of a single hydrogen bond between a carboxylate oxygen and N7 of the purine ring. These results and others discussed below are in agreement with the postulated role of the adjacent aspartates in the catalytic mechanism for mADA.
- Department of Biochemistry and Cell Biology, Rice University, Houston, Texas 77005, USA.
Organizational Affiliation: