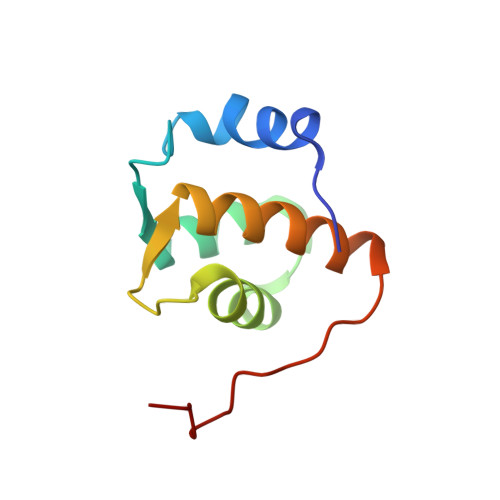
Solution structure of the Reps1 EH domain and characterization of its binding to NPF target sequences.
Kim, S., Cullis, D.N., Feig, L.A., Baleja, J.D.(2001) Biochemistry 40: 6776-6785
- PubMed: 11389591 Search on PubMed
- DOI: https://doi.org/10.1021/bi002700m
- Primary Citation Related Structures:
1FI6 - PubMed Abstract:
The recently described EH domain recognizes proteins containing Asn-Pro-Phe (NPF) sequences. Using nuclear magnetic resonance (NMR) data, we determined the solution structure of the EH domain from the Reps1 protein and characterized its binding to linear and cyclic peptides derived from a novel targeting protein. The structure calculation included 1143 distance restraints and 122 angle restraints and resulted in structures with a root-mean-square deviation of 0.40 +/- 0.05 A for backbone atoms of superimposed secondary structural elements. The structure comprises two helix-loop-helix motifs characteristic of EF-hand domains. Titration data with NPF-containing peptides showed evidence of intermediate exchange on the NMR chemical shift time scale, which required an analysis that includes curve fitting to obtain accurate equilibrium constants and dissociation rate constants. The cyclic and linear peptides bound with similar affinities (Kd = 65 +/- 17 and 46 +/- 14 microM, respectively) and to the same hydrophobic pocket formed between helices B and C. The cyclic peptide formed a complex that dissociated more slowly (k(off) = 440 +/- 110 s(-1)) than the linear peptide (k(off) = 1800 +/- 250 s(-1)), but had little change in affinity because of the slower rate of association of the cyclic peptide. In addition, we characterized binding to a peptide containing a DPF sequence (Kd = 0.5 +/- 0.2 mM). The characterization of binding between the Reps1 EH domain and its target proteins provides information about their role in endocytosis.
- Department of Biochemistry, Tufts University School of Medicine, 136 Harrison Avenue, Boston, Massachusetts 02111, USA.
Organizational Affiliation: