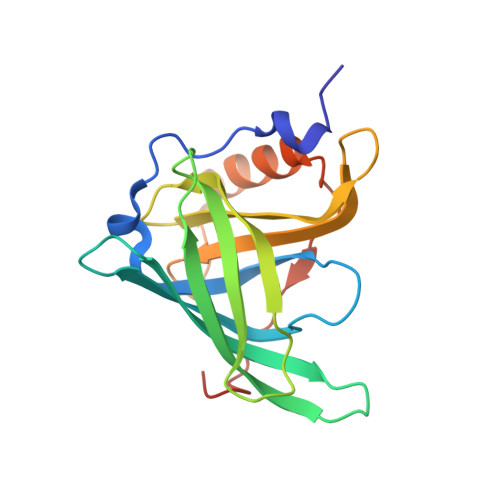
Crystallographic studies on complexes between retinoids and plasma retinol-binding protein.
Zanotti, G., Marcello, M., Malpeli, G., Folli, C., Sartori, G., Berni, R.(1994) J Biological Chem 269: 29613-29620
- PubMed: 7961949 Search on PubMed
- Primary Citation Related Structures:
1FEL, 1FEM, 1FEN - PubMed Abstract:
The three-dimensional structures of complexes between bovine plasma retinol-binding protein (RBP) and three retinol analogs with different end groups (fenretinide, all-trans retinoic acid, and axerophthene) have been determined to 1.8-1.9-A resolution. Their models are very similar to that of the bovine retinol.RBP complex: the root mean square deviations between equivalent alpha-carbons in the two proteins range from 0.17 to 0.24 A. The retinoid molecules fit in the beta-barrel cavity assuming the same conformation of the vitamin, and the substitutions have no consequences on the overall protein structure. While confirming that an intact hydroxyl end group is not an absolute requirement for a correct retinoid binding to RBP, this study has shown the occurrence of conformational changes, although limited, in the rather flexible loop region at the entrance of the beta-barrel upon fenretinide and retinoic acid binding. These changes are suitable for accommodating the end groups of the above retinoids. Instead, no such changes have been revealed in RBP complexed with axerophthene, a retinol analog bearing a hydrogen atom in place of the hydroxyl end group. The protein conformational changes in the above loop region, the steric hindrance of bulky end groups of bound retinoids, and the lack of the retinol hydroxyl group appear to be responsible for the possible reduced affinity of retinoids for RBP relative to retinol and, at the same time, for the abolished or reduced affinity of retinoid.RBP complexes for transthyretin relative to retinol-RBP.
- Department of Organic Chemistry, University of Padova, Italy.
Organizational Affiliation: